Summary
MEMBRANE PROTEIN STRUCTURAL BIOLOGY
The lab is based at the Research Complex at Harwell, Oxfordshire (https://www.rc-harwell.ac.uk/research-groups/beis-group/).
Membrane proteins represent around 30% of the proteomes of most organisms and more than 40% of drug targets and yet few structures of these molecules have been solved by x-ray crystallography. Drug resistance of bacterial pathogens is a rising crisis. Bacterial membrane proteins are essential for resistance since they are involved in the export of the drugs from the cell. My group is interested to determine the structure and function of these proteins in order to elucidate the molecular mechanism of drug resistance.
The lab is also interested to exploit novel antibacterials as treatments for bacterial infections by understanding how bacteria secrete antibacterial peptides and hijack membrane transporters of other bacteria to inhibit their growth under nutrient starvation.
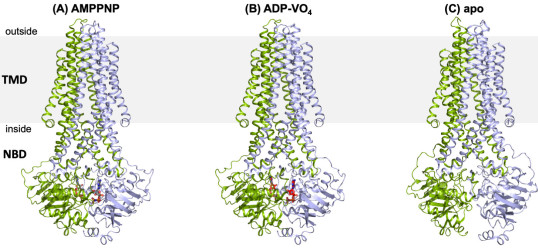
We are also investigating the mechanism and dynamics of ABC transporters by employing a multidisplinary approach of protein crystallography, transport assays and biophysical methods such as PELDOR/EPR and smFRET.
Publications
Journals
Griffiths D, Anderson M, Richardson K, et al., 2024, Cyclic Ion Mobility for Hydrogen/Deuterium Exchange-Mass Spectrometry Applications., Anal Chem, Vol:96, Pages:5869-5877
Baquero F, Beis K, Craik DJ, et al., 2024, The pearl jubilee of microcin J25: thirty years of research on an exceptional lasso peptide., Nat Prod Rep, Vol:41, Pages:469-511
Smith HE, Mackenzie AM, Seddon C, et al., 2024, The use of NADH anisotropy to investigate mitochondrial cristae alignment., Sci Rep, Vol:14
Mazza T, Roumeliotis TI, Garitta E, et al., 2024, Structural basis for the modulation of MRP2 activity by phosphorylation and drugs, Nature Communications, Vol:15, ISSN:2041-1723
Seddon C, Frankel G, Beis K, 2024, Structure of the outer membrane porin OmpW from the pervasive pathogen Klebsiella pneumoniae, Acta Crystallographica Section F: Structural Biology and Crystallization Communications Online, Vol:80, ISSN:1744-3091, Pages:22-27

