Overview
Structural biology of extracellular matrix
Extracellular matrix (ECM) is assembled from secreted macromolecules by a combination of spontaneous and cell-assisted processes. Cells interact with ECM using membrane-spanning receptors that regulate cell behaviour. Many human diseases are the result of faults in ECM assembly or cell-matrix communication. For example, defective adhesion of skeletal muscle cells to ECM causes muscular dystrophy, and the loss of regulation by ECM is a key step in the progression of many cancers.
The aim of my research is to understand ECM assembly and signalling at the molecular level.
Laminins
The heterotrimeric laminins are a major constituent of basement membranes, which are a type of ECM that underlies all epithelia and surrounds muscle cells. Laminin polymerisation is mediated by the LN domains at the tips of laminin's three short arms. We determined the structure of the LN domains and, in collaboration with Peter Yurchenco (Rutgers University), showed how they interact to form the nodes of the laminin polymer (Hussain et al, 2011; Purvis and Hohenester, 2012; McKee et al, 2021).
The laminin polymer is anchored to the cell surface by interactions of the LG domains with cellular receptors, such as integrins and dystroglycan. In 1999, we proposed that LG domains may function as calcium-dependent lectins for the then uncharacterised carbohydrate modification of dystroglycan (Hohenester et al, 1999). In collaboration with Kevin Campbell (University of Iowa), we subsequently confirmed this prediction by elucidating the atomic details of this important protein-carbohydrate interaction (Briggs et al, 2016).
A long standing question has been why integrins recognise only laminin heterotrimers, and not any of the single chains. Our structure of a heterotrimeric laminin-111 fragment revealed how the α1, β1 and γ1 chains combine to form a functional integrin binding site (Pulido et al, 2017).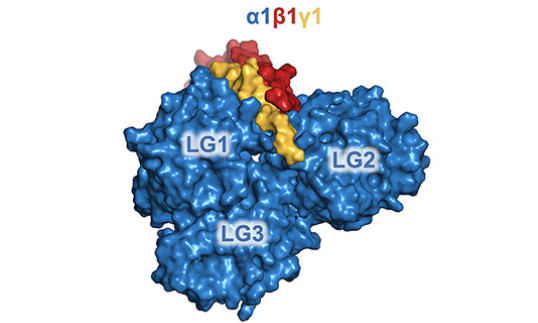 Collagens
Collagens
Collagens are the major structural proteins of vertebrates, but they also serve as ligands of two unusual receptor tyrosine kinases, DDR1 and DDR2. In collaboration with Birgit Leitinger (National Heart and Lung Institute), we defined the structural basis of collagen recognition and signalling by DDRs (Carafoli et al, 2009; Carafoli et al, 2012; Sammon et al, 2020).
The degradation of collagens is carried out by matrix metalloproteinases (MMPs). In collaboration with Hideaki Nagase (Kennedy Institute of Rheumatology) and Richard Farndale (University of Cambridge), we determined the long-elusive structure of MMP-1 bound to a triple-helical collagen-like peptide, which revealed how the enzyme is targeted to the unique cleavage site in fibrillar collagens (Manka et al, 2012).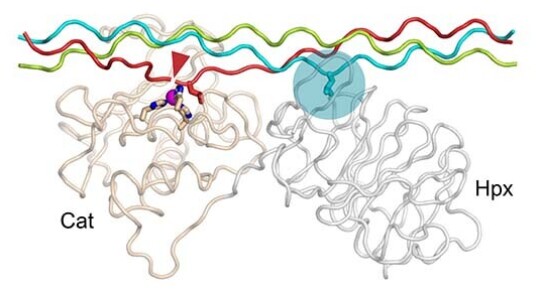 Glycosaminoglycan biosynthesis
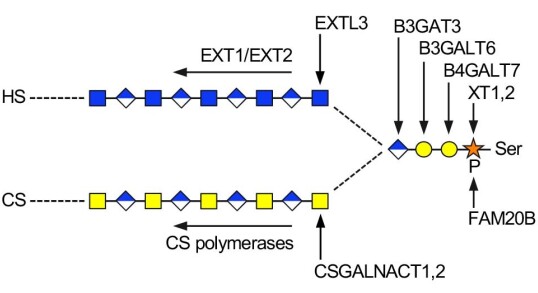
Glycosaminoglycan biosynthesis
Ηeparan sulfate (HS) and chondroitin sulfate (CS) are protein-attached glycosaminoglycan chains that play critical roles in animal development and physiology. How the right chain type (HS or CS) is attached to a given core protein has been an enduring mystery. We are tackling this question by reconstituting glycosaminoglycan biosynthesis in vitro. We discovered how the first enzyme in the pathway, xylosyltransferase, selects serine residues for modification (Briggs and Hohenester, 2018). More recently, we established how the branching of the pathway is controlled: CSGALNACTs are capable of initiating CS synthesis on all core proteins, whereas EXTL3 must recognise specific features in the core protein to initiate HS synthesis (Sammon et al, 2023).