Bacterial-mediated regulation of host cell signalling
When mounting an immune response against pathogens, host cells undergo significant changes in gene expression. A key aim of the laboratory is to understand how bacterial virulence proteins alter this response, dampening host immunity and promoting the pathogen. We therefore use a combination of biochemistry, proteomics and CRISPR technology to investigate the points of intersection between host cell signalling and pathogen virulence factors.
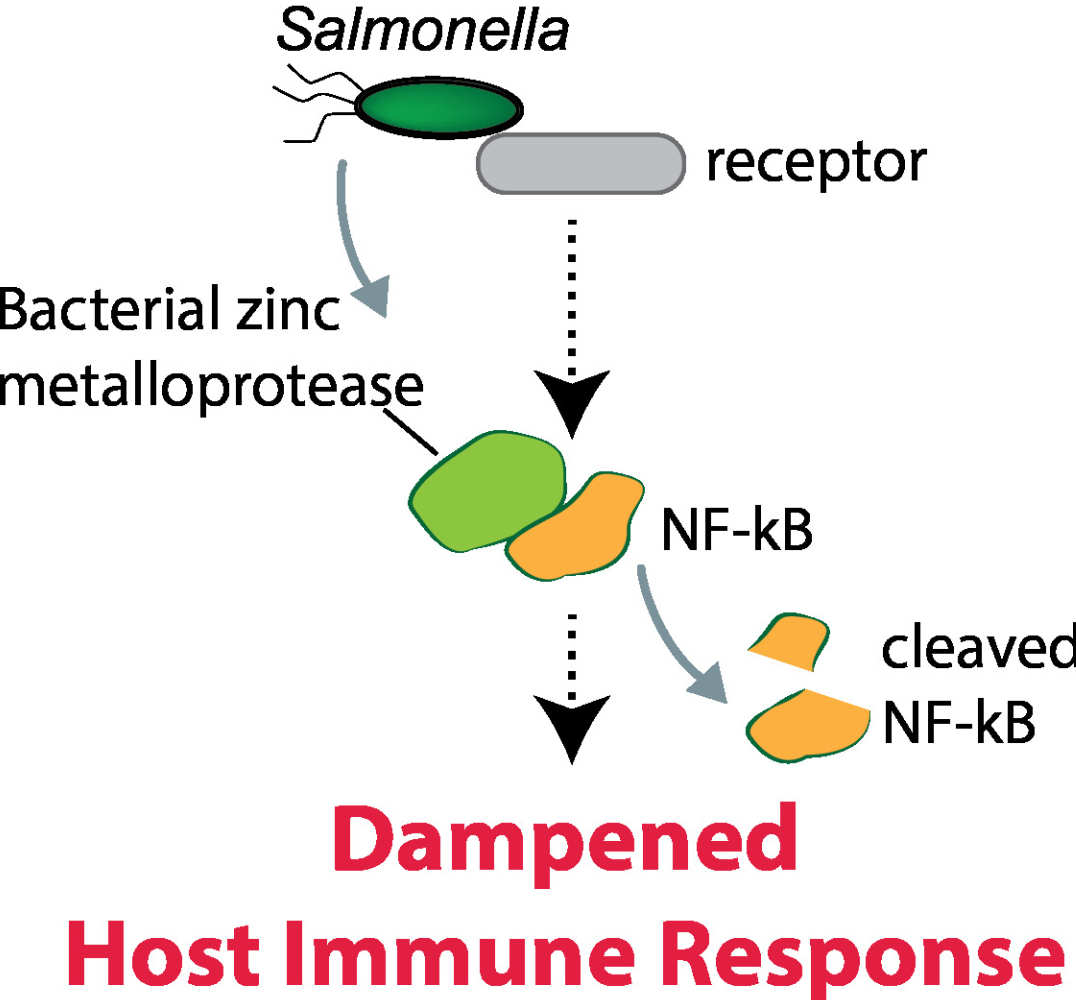
Activity of the Salmonella effector GtgA
Host responses to bacterial pathogens

TRIM proteins represent a family of E3 ubiquitin ligases with a variety of functions in the resistance to pathogens, including pathogen recognition and the regulation of immune signal transduction and transcription factor activation. Our aim is to further our understanding on the role of TRIM family members during infection of immune cells by intracellular pathogens.
Techniques

We employ a broad range of techniques to explore the protein-protein interactions that govern the outcome of bacterial infections. These include immuno-precipitation and mass spectrometry experiments to identify novel protein interactions, proteomics to explore protein post-translational modifications and reporter-based assays to investigate novel functions of bacterial proteins. We combine this with the powerful technique of CRISPR to generate gene knockouts, studying which genes are required for host immunity and the restriction of bacterial pathogens.
Immuno-fluorescence microscopy
Seeing is believing. We use fixed and live cell imaging to visualise bacterial infections at the single cell level and over time.
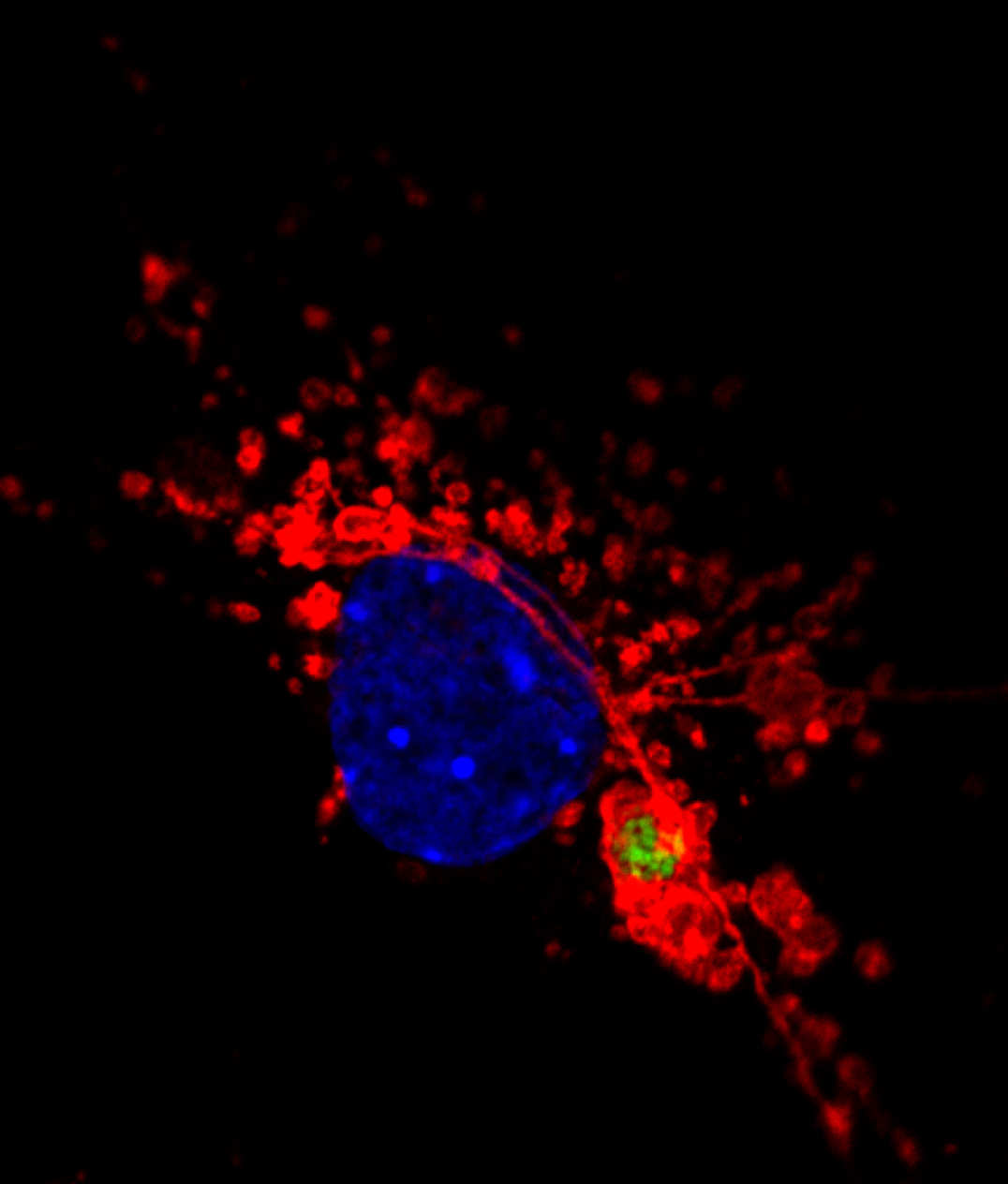
Salmonella infected MEF cell. Lamp1-red, Salmonella-green, DNA-blue
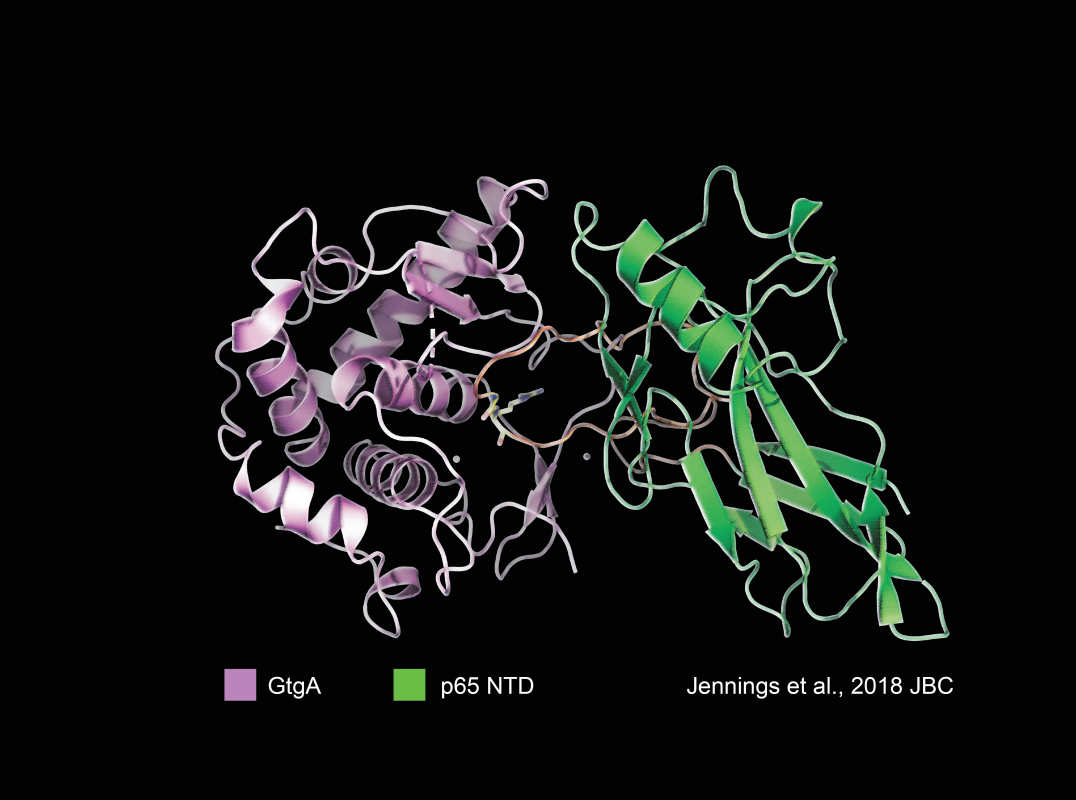
The bacterial effector GtgA in complex with its host target, p65
Structural studies
In collaboration with the Rittinger Lab at the Crick Institute, we employ X-ray crystallography in order to understand the molecular determinants of bacterial effector protein function and their interactions with host targets.
Collaborators
Prof. Denise Monack, Stanford University, Salmonella persistence in vivo, 2018
Prof. Steven Ley, Imperial College London, Innate immune signaling, 2017
Katrin Rittinger, The Crick Institute, Molecular Structure of Cell Signalling, 2016
Guest Lectures
Innate Immune Inhibition by Salmonella effector proteins, Crick - Dundee signalling symposium, The Crick Institute, 2017
Galectin-8 catches cytosol-invading Salmonella, EMBO workshop, Mandelieu-la-Napoule, France, 2016
Host pathogen interactions and cell-autonomous immunity, University of Dundee, Dundee, 2014
Research Student Supervision
Guenster,R, Functional analysis of the SseK family of Salmonella SPI-2 type III secretion system effectors (PhD)
Jennings,E, Salmonella-mediated inhibition of innate immunity (PhD)
Ong,S, Regulation of AP1 by Salmonella
Panagi,I, Salmonella effector function and immune signaling
Tedros,Y, Investigating Trim32 interactions partners
Wong,S, Monitoring cytosolic Salmonella

