Welcome to the Genomics Core Facility!
The Genomics Core Facility, provides state-of-the-art genomics and proteomics equipment and services to researchers within the Department of Bioenigneering, collaborators and from time to time external customers.
The facility has close ties to many others including, Microbiology (e.g. for cloning), in-vivo (genotyping and expression studies) and well as cell and tissue culture analysis.
Genomics info
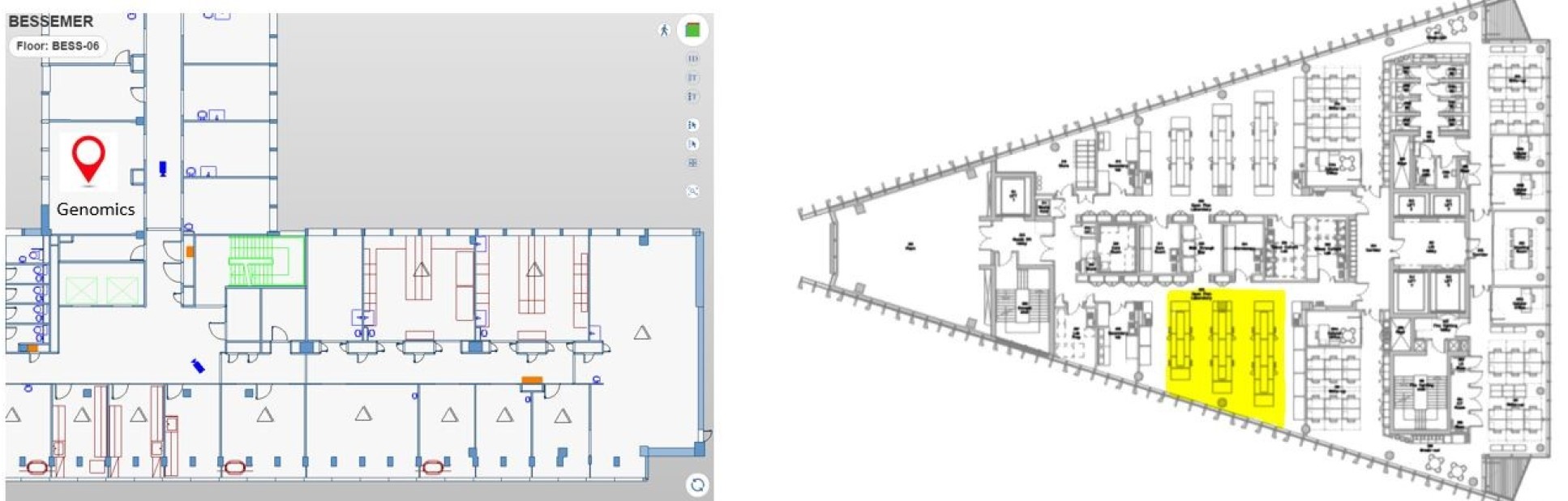
Room B627 Bessemer Building, South Kensington Campus Room 302, Sir Michael Uren Hub, White City Campus
See risk assessments: Risk assessment | Faculty of Engineering | Imperial College London
GenomicsItems
- Access: Induction and Training
- Aseptic Techniques
- Equipment
- Induction
- Overview
- PCR and QPCR
- Plate reader
Access to the facility is granted by the core facility manager after completing the Health and Safety induction and equipment training. The completion of a Risk Assessment is comulsory before starting the work in the Facility.
To request access after completing the required training and room introduction, sign and upload the induction form and complete the access request form on the following link: https://www.imperial.ac.uk/bioengineering/admin/info/swipe/

 | ||
|---|---|---|
| PCR (Polymerase Chain Reaction) Tutorial - An Introduction | 1. Pdf manual – different types of PCR and their application | 1. Standard Protocol |
| 2. Primer design basics | 2. Intro to Techniques - Springer | |
| QPCR analysis |
3. Avoiding contamination | |
| Enter table footer here | ||
Successful cell culture depends heavily on keeping the cells free from contamination by microorganisms such as bacterial, fungi, and viruses. Nonsterile supplies, media, and reagents, airborne particles laden with microorganisms, unclean incubators, and dirty work surfaces are all sources of biological contamination.
Aseptic technique, designed to provide a barrier between the microrganisms in the environment and the sterile cell culture, depends upon a set of procedures to reduce the probability of contamination from these sources. The elements of aseptic technique are a sterile work area, good personal hygiene, sterile reagents and media, and sterile handling.
Essential Sterile Technique
The steps to prevent contamination of your cell culture and demonstration of best-practice sterile techniques.
Sterile work area
The simplest and most economical way to reduce contamination from airborne particles and aerosols (e.g., dust, spores, shed skin, sneezing) is to use a cell culture hood.
- The cell culture hood should be properly set up and be located in an area that is restricted to cell culture that is free from drafts from doors, windows, and other equipment, and with no through traffic.
- The work surface should be uncluttered and contain only items required for a particular procedure; it should not be used as a storage area.
- Before and after use, the work surface should be disinfected thoroughly, and the surrounding areas and equipment should be cleaned routinely.
- For routine cleaning, wipe the work surface with 70% ethanol before and during work, especially after any spillage.
- You may use ultraviolet light to sterilize the air and exposed work surfaces in the cell culture hood between uses.
- Using a Bunsen burner for flaming is not necessary nor is it recommended in a cell culture hood.
- Leave the cell culture hood running at all times, turning it off only when they will not be used for extended periods of time.
Good personal hygiene
Wash your hands before and after working with cell cultures.
In addition to protecting you from hazardous materials, wearing personal protective equipment also reduces the probability of contamination from shed skin as well as dirt and dust from your clothes.
Sterile reagents & media
Commercial reagents and media undergo strict quality control to ensure their sterility, but they can become contaminated while handling. Follow the guidelines below for sterile handling to avoid contaminating them. Always sterilize any reagents, media, or solutions prepared in the laboratory using the appropriate sterilization procedure (e.g., autoclave, sterile filter).
- Always wipe your hands and your work area with 70% ethanol.
- Wipe the outside of the containers, flasks, plates, and dishes with 70% ethanol before placing them in the cell culture hood.
- Avoid pouring media and reagents directly from bottles or flasks.
- Use sterile glass or disposable plastic pipettes and a pipettor to work with liquids, and use each pipette only once to avoid cross contamination. Do not unwrap sterile pipettes until they are to be used. Keep your pipettes at your work area.
- Always cap the bottles and flasks after use and seal multi-well plates with tape or place them in resealable bags to prevent microorganisms and airborn contaminants from gaining entry.
- Never uncover a sterile flask, bottle, petri dish, etc. until the instant you are ready to use it and never leave it open to the environment. Return the cover as soon as you are finished.
- If you remove a cap or cover, and have to put it down on the work surface, place the cap with opening facing down.
- Use only sterile glassware and other equipment.
- Be careful not to talk, sing, or whistle when you are performing sterile procedures.
- Perform your experiments as rapidly as possible to minimize contamination.
-
To note, usage charges cover maintenance and running of equipment. New equipment is added by academic cost sharing and grant proposals.
Departmental Equipment Wish List - Items of equipment can be added to the departmental equipment wish list for addition to future PI grants and shared multi-user grant proposals. Examples of current items include a new cryostat, microtome and embedding station. Once added to the core facility the equipment is management, serviced and maintained by the Core Facility Technical Team.
South Kensington

Labnet's MultiGene Gradient Thermocycler
This thermocycler provides consistent, reliable results while making gradient technology affordable. This unit is extremely simple to program, compact in design and built to perform.
Step One Plus Real-Time PCR machine
A 96 well instrument for applications such as SNP Genotyping, Gene Expression Analysis, MicroRNA Expression, Protein Expression, Translocation Analysis, Gene Detection, Viral Load Analysis.

A wide choice of electrophoresis apparatus and blotting apparatus is provided for protein and nucleic acid applications

BD Accuri C6 Flow Cytometer with sampler
A unique low-pressure pumping system drives the fluidics. A sheath-focused core enables event rates of up to 10,000 events per second and a sample concentration of over 5 x 106 cells per mL. In addition, the system derives sample volume and can calculate absolute counts or sample concentration per microliter.

UVP Sterile PCR and workstationEquipped with an open UV lamp installed in the upper part of the hood. Ideal for Nucleic acid extractions, PCR.

UVP Biospectrum imaging system
Gel doc, DNA, protein gels, chemiluminescence blots, TLC gels and 2D gels with 5MP camera)

ThermoFisher Fluroskan FL Microplate Fluorometer and Luminometer
Perform a number of fluorometric and luminometric research applications with the Thermo Scientific™ Fluoroskan Ascent™ FL Microplate Fluorometer and Luminometer, which offers versatile plate formats, fast plate reading speeds, top/bottom plate reading and up to three dispensers

Thermo Scientific Varioskan Flash Plate readerDelivers excellent performance in imperious research assays and spectral scanning. This equipment ensures remarkable versatility for a huge range of implementations with the inclusion of up to three onboard dispensers, unconditional wavelength selection, effortless measurement setup with automated internal functions, incomparable optical performance in addition to a highly innovative Thermo Scientific SkanIt* Software.

Eppendorf Biospectrophotometer
Measurement of DNA, RNA and protein concentration (UV and colourimetric). Applications: Incorporation rate of fluorescent molecules (550 nm/650nm), e.g. for microarray experiments, Enzymatic assays (e.g., peroxidase, alkaline phosphatase, beta-galactosidase) and Optical density of cells (OD 600) etc.

1.5 and 2ml tubes. 13,000rpm.

Departmental Equipment Wish List
Items of equipment can be added to the departmental equipment wish list for addition to future PI grants and shared multi-user grant proposals. Examples of current items include a new cryostat, microtome and embedding station. Once added to the core facility the equipment is management, serviced and maintained by the Core Facility Technical Team.
White City

Clariostar Plus
The Clariostar Plus plate reader is a high-performance microplate reader offering a range of application including UV/vis absorbance, protein-protein interactions, enzyme activity and kinetic assays and cell-based assay. The LVis plate that comes with this reader can quantify concentrations of DNA and RNA in up to 12 samples.

QPCR Quantstudio 3
The QuantStudio is a simple real-time PCR machine which allows users to look at gene expression, copy number variation, protein detection and various other applications. With this software users can remotely monitor runs, results and collaborate with colleagues.

can a ThermoFisher Proflex 3X32 Thermocycler
This ProFlex Thermocycler has 3 individual blocks containing 32 wells each allowing users to run three separate experiments simultaneously, with an easy to use touchscreen interface.

Azure 280
The Azure 280 is a high-performance imaging system with an easy to use interface allowing the user to take quick pictures of gels and blots. The 280 features colour imaging, blue and UV light imaging and chemiluminescence.

Sonicator Q500
The Q500 is an easy to use sonicator which uses ultrasonic sound to process samples. It has applications such as nanoparticle dispersion, cell lysis, homogenization and much more and can be programmed for up to 10 hours. This sonicator has a sound enclosure to protect the user from the high pitch noise and reducing the noise by approximately 20 dBa.

Electrophoresis equipment
A range of equipment from Biorad such as gel tanks, combs, power supplies, docking stations, etc to run agarose gels and Western blots.
Summary of the table's contents
|
User Registration
Survey Powered By Qualtrics
|
The Genomics Facility, is a multi-user laboratory managed and overseen by the departmental dedicated technician \ facility manager (with support from the wider technical team), in the Department of Bioengineering to provide core Genomics and proteomics equipment and services, including
- PCR (including RT, Q as well as standard PCR)
- UV cabinets for sterility
- Gel electrophoresis and imaging
- Plate readers
All researchers using this space must adhere to strict rules on aseptic techniques, including housekeeping standards of care and quality. This is both to protect the users, but is essential for protecting the work and preventing contamination of genomic samples.
PCR can be decribed in 3 broad terms, each with their own use and requirements
- PCR - Polimerase Chain reaction - a basic check of the presence of genes, often used for genotyping
- QPCR - quantatative PCR - a quantification of how many of a gene is present in the sample
- RT-PCR - Reverse transcriptase PCR - Analysis of RNA's and genomic transcriptions
1. PCR
PCR indu
ction
2. Q-PCR
QPCR
3.
RT-qPCR

varioskan
The Thermo Scientific Fluoroskan FL is a compact and robust microplate fluorometer and luminometer, offering simplicity and reliability for increased efficiency of workflows. Featuring excellent optical performance, the Fluoroskan FL Microplate Fluorometer and Luminometer is ideal for life science fluorometric and luminometric assays. The Fluoroskan FL fluorometer and luminometer offers plate format versatility, fast plate reading speeds, and up to three dispensers.
• High sensitivity for both top and bottom reading
• Fast reading speed essential for kinetic applications
• Onboard dispensers for precise initiation of kinetic reactions
• Powerful and easy-to-use Thermo Scientific SkanIt software
High sensitivity for both top and bottom reading
The use of fiberless direct illumination optics for both top and bottom reading enables high sensitivity, a wide dynamic range, and accurate and precise results for 6- to 384-well plates.
Fast reading speed essential for kinetic applications
The Fluoroskan FL Microplate Fluorometer and Luminometer is capable of reading a 96-well plate in just 18 seconds, essential for kinetic applications. Fluorometric GFP measurement and luminometric luciferase measurement from the same well occurs quickly due to the versatility of the assay programming.
Onboard dispensers precise initiation of kinetic reactions
The Fluoroskan FL Microplate Fluorometer and Luminometer can be equipped with up to three reagent dispensers* for use during fast kinetic assays, such as Ca2+ flux assays. The onboard dispensers support simultaneous dispensing and reading, enabling the monitoring of fast kinetic measurements from the very start of the reaction. Very low dead volume and backflush capability support conservation of reagents.
*Upgrade to third dispenser must be performed by service technician
Powerful and easy-to-use SkanIt software
With a visual workflow setup, SkanIt software is quick and easy to learn. It offers effortless data analysis (with a comprehensive selection of built-in calculations, including blank subtraction, standard curve creation, classification, quality control, kinetic analysis) and exporting capabilities. Each user can install SkanIt software on his/her own computer to set up assays and analyze data.
Includes:
SkanIt PC Software and the following filter pairs (excitation/emission): 355 nm/460 nm, 485 nm/538 nm. An extensive selection of additional filters is available upon request.
Applications include:
Ca2+ flux assays , cell proliferation, cytotoxicity, cell adhesion, nucleic acid quantitation, reporter gene assays, hybridization assays, ion channel, FRET assays, BRET assays, immunoassays, enzyme activity, bacterial quantitation, phagocytosis, oligonucleotide assays









