The Imperial researchers taking the fight to the superbugs
Carbapenem-resistant enterobacteriaceae (CRE)
A new collaborative aims to curb the growing threat of antimicrobial resistance with prescribing tools, digital infection diagnosis & new antibiotics
It’s very difficult to imagine life before the era of antibiotics. Aggressive infections such as tuberculosis, pneumonia and whooping cough were common killers – and it was not unheard of for minor wounds and burns to become infected and prove fatal.
 That all changed with the discovery of penicillin by Sir Alexander Fleming (left) at St Mary’s Hospital in 1928 and its subsequent therapeutic development in the early 1940s involving another Imperial scientist, Sir Ernst Chain.
That all changed with the discovery of penicillin by Sir Alexander Fleming (left) at St Mary’s Hospital in 1928 and its subsequent therapeutic development in the early 1940s involving another Imperial scientist, Sir Ernst Chain.
The ensuing decades brought the development of a raft of highly effective new antibiotics, and in 1969 US Surgeon General, William Stewart, boldly told Congress it was time to ‘close the books on infectious diseases’. Yet, just as Fleming had presciently warned, bacterial strains began to emerge that had acquired resistance to these very antibiotics.
The lack of new drug development in the 1980s and 1990s, coupled with an explosion in the use of existing antibiotics in both humans and farm animals has hugely escalated the resistance problem – to the point where some experts now believe we are entering a ‘post-antibiotics era’.
In 2014 a UK government review estimated that failing to tackle antimicrobial resistance (AMR) would result in at least 10 million extra deaths a year by 2050 (more than the number of people who currently die from cancer), and Prime Minister David Cameron promised to make the issue a national priority.
On many fronts
Imperial plans to play just as pivotal a role in the fight back against antimicrobial resistance as it did in the development of the first antibiotics.
The new Antimicrobial Research Collaborative (ARC) is a network of researchers, clinicians and healthcare professionals from Imperial’s four faculties, Imperial College Healthcare NHS Trust, and industry who have joined forces to address the global threat of AMR.
ARC is led by Professor Alison Holmes (pictured right), Professor of Infectious Diseases in the Department of Medicine and Director of Infection Prevention and Control for the Trust.
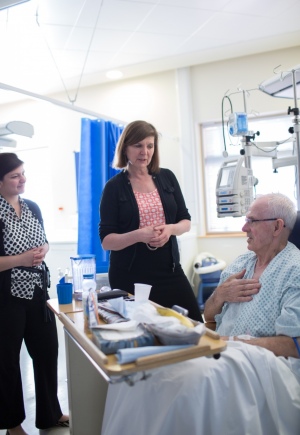
Alison Holmes at the General Adult Intensive Care Unit at Hammersmith Hospital
“The rapid spread of AMR is a serious global public health issue,” she says. “If we don’t address this, infection which were once easily treatable will become fatal.
“ARC will tackle this problem by adopting a collaborative approach across a broad range of disciplines.”
This will encompass structural biology right through to health policy initiatives and employ the latest techniques including big data, microelectronics and synthetic biology. The end goal will be the same though: to translate research for the benefit of patients as quickly as possible.
“This is not just about the treatment of infections here and now, for you and me − it’s also for the next generation,” Alison adds. “It is vitally important not just from a healthcare perspective but also from a societal perspective.”
We spoke with some of those working as part of the ARC.
Empowering doctors
ARC collaborator Dr Luke Moore is a Registrar in Infectious Diseases at Imperial College Healthcare NHS Trust and Clinical Research Fellow in the Department of Medicine, working in Alison Holmes’ group. Luke practiced as a medical doctor for ten years, trying out different areas of medicine including cardiology. But after witnessing the AMR threat first hand was galvanised to play a part in the remedy.

Luke Moore at Imperial Festival
“Ninety-five per cent of my clinical work involves complex antibiotic choices, because the bugs have become resistant to the most commonly used drugs,” he says. “Outbreaks are common, especially in London. Right now, we’ve been dealing with a strain that is resistant to every single drug bar one, called colistin, which was developed in the 1960s but largely abandoned because it can have a significant impact on kidney function. But our backs are against the wall; it’s a last resort.”
Luke and others believe the first step is to address one of the root causes of the problem which has led us to this position – an explosion in antibiotic use, often with the ‘blunt instrument’ of broad spectrum antibiotics.
“We know about Darwinian pressure, we know how bugs become resistant; yet still, up to half of all antibiotics prescribed are inappropriate. So every other time a doctor puts pen to paper, it may not have the desired outcome and will likely exacerbate resistance.”
 Luke is working with computing experts to develop ‘machine learning’ decision-support tools for hospital doctors who routinely prescribe antibiotics. Delivered on ipads and mobile phones, the applications use available information on the patient and their infection – but crucially also pull in data from the cloud on similar patients in the locality. This means the app is constantly updating with data from the latest outbreaks and is able to ‘learn’ from this when advising the doctor.
Luke is working with computing experts to develop ‘machine learning’ decision-support tools for hospital doctors who routinely prescribe antibiotics. Delivered on ipads and mobile phones, the applications use available information on the patient and their infection – but crucially also pull in data from the cloud on similar patients in the locality. This means the app is constantly updating with data from the latest outbreaks and is able to ‘learn’ from this when advising the doctor.
The app is currently being trialled in hospitals and Luke and the team hope to develop and roll out a version for GPs at some point in the future.
Rapid diagnosis
Prescribing tools have the potential to become even more powerful when coupled with rapid diagnosis of the underlying infection. For several years, Regius Professor Christofer Toumazou (Bioengineering) and his team has been developing microchip technology that directly translates DNA code into digital information for various patient healthcare applications.
However, he soon saw the potential of the technology to reveal the identity of culprit bacteria responsible for infections, just as forensics identifies a perpetrator. This is particularly important in the management of bloodstream infections and sepsis – a major worldwide killer. When sepsis is suspected, clinicians urgently need to know the type of infection before administering the appropriate type of antibiotics.
However, this usually requires remote laboratory analysis which can take days to complete, by which time, the patient’s condition can become much worse. With little information doctors often prescribe inappropriate antibiotics, to the detriment of the patients and potentially fuelling the resistance problem.
 “My son has relied on renal dialysis for many years now, so I am quite familiar with the acute hospital environmental − and frankly it is primitive with respect to adoption of the latest technologies,”says Chris. “Only recently he had a potential infection scare in hospital, which luckily turned out to be negative, but it was agonising waiting for the blood cultures to be analysed. Fortunately though, we have a better way, and we believe it will be in the clinic very soon.”
“My son has relied on renal dialysis for many years now, so I am quite familiar with the acute hospital environmental − and frankly it is primitive with respect to adoption of the latest technologies,”says Chris. “Only recently he had a potential infection scare in hospital, which luckily turned out to be negative, but it was agonising waiting for the blood cultures to be analysed. Fortunately though, we have a better way, and we believe it will be in the clinic very soon.”
Chris’ prototype device looks like a slightly larger USB pen drive, and works in a similar fashion, taking samples at one end and uploading the data to a laptop at the other, allowing identification of the infection in as little as a couple of hours instead of days.
The next generation
Whilst better use of existing drugs will help to curb the resistance problem, many experts believe that in the long-term we will ultimately need an entirely new repertoire of treatments. Many researchers in the ARC are working on just that, honing in on different mechanisms of attack, some that work in a similar fashion to existing antibiotics while others that attempt to find new bacterial vulnerabilities.
One group is attempting to use the power of synthetic biology − where cells are genetically reprogrammed as production factories − to explore the possibilities of manufacturing new antibiotics.
 ARC collaborator Dr Tom Ellis (Bioengineering; left), explains that the first step in this ambitious project is to establish what he refers to as ‘warfare system’, where yeast cells are reprogrammed to produce penicillin, then grown in the presence of a bacterial cocktail.
ARC collaborator Dr Tom Ellis (Bioengineering; left), explains that the first step in this ambitious project is to establish what he refers to as ‘warfare system’, where yeast cells are reprogrammed to produce penicillin, then grown in the presence of a bacterial cocktail.
“These synthetic yeast cells will only grow if they can produce sufficient yields of penicillin. So we will set up an array of 1,000 little tubes and add new bits of DNA that we suspect may give an advantage – for example by increasing or controlling the amount of penicillin production.”
Once this system is established they can start to design entirely new antibiotic structures, and select for those that appear to be effective against resistant bacterial strains, using their yeast cultures for confirmation.
“Because many enzymes that produce antibiotics tend to be modular themselves, it really lends itself to the synthetic biology approach, where we can experiment with different combinations. Ultimately we could have a system that sifts through billions of combinations − we call it the broth of possibilities!”
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Andrew Czyzewski
Communications Division