Study sheds light on body's sticky "sugar forests" where viruses take hold
Researchers have pioneered a new method for pinpointing the sugar molecules in mucus where intestinal viruses "stick" to and infect cells.
In the study, published in Molecular & Cellular Proteomics, the team used the technique, dubbed ‘Beam Search’, to zoom in on the sticky forest-like chains of carbohydrates at the surface of our intestines where microbes take hold.
The research, led by Professor Ten Feizi in the Department of Medicine, could in future lead to more intelligent vaccine designs and new treatments for common rotaviruses and noroviruses.
Gateways to infection
The cells covering the surface of our intestine are densely decorated with intricate carbohydrate chains known as glycans. These glycans consist of strings of sugar molecules which, when viewed through an electron microscope, look something like a forest. They are often referred to as a glycocalyx.
Viruses are covered with adhesive proteins that seek out and select specific glycans to stick to. This enables them to get a foothold on the particular cells that they are trying to infect. Rotaviruses, for instance, will recognise specific glycans that cover the cells on our intestine’s surface.
These so-called glycan forests are particularly dense where there are mucins – high molecular weight mucous molecules that sit on the surface of certain cells. It is in these areas that Professor Feizi’s team chose to focus their new approach, with the aim of pinpointing the glycans being selectively bound by the adhesive proteins of two different rotaviruses.
Finding the sticking point
We anticipate that similar studies will help us to unravel the relationship of the blood groups of individuals to their susceptibilities to other viruses that cause gastroenteritis Professor Ten Feizi
Using existing techniques, researchers are required to carry out lengthy separation procedures to isolate all glycans present on a particular molecule, before carrying out systematic testing to uncover the glycan bound by the virus.
However, when using Beam Search, the Imperial team were able to quickly focus on the bioactive components recognised by the virus. Having zoned in on a particular mucin, several hundred glycans were released, fractionated and robotically arrayed in order to focus on the fraction where the rotaviruses would stick. After one additional fractionation step and arraying, the researchers were able to purify and determine the sequence that had been selectively bound by the viral protein.
Significantly, the study found that the two rotaviruses tested had bound to a long glycan sequence, containing the blood group O antigen.
Developing better vaccines
Commenting on the implications of the study, Professor Feizi said: “The power of the new approach is analogous to finding a needle in a haystack!”
“We anticipate that similar studies will help us to unravel the relationship of the blood groups of individuals to their susceptibilities to other viruses that cause gastroenteritis. This will inform live vaccine designs and provide lead compound for therapeutic designs.”
“However, although the Beam Search approach is orders of magnitude more sensitive than traditional methods of glycan ligand discovery, there is scope for miniaturization of the requirement of starting materials, and there are plans in hand for this.”
‘O-Glycome Beam Search Arrays for Carbohydrate Ligand Discovery’ by Zhen Li, Chao Gao et al. is published in Molecular & Cellular Proteomics.
Find out more about ongoing research at Imperial's Glycosciences Laboratory.
The study was funded by the Wellcome Trust, and was supported in part by the China Scholarship Council.
The adhesive proteins of the rotaviruses were prepared in the collaborating laboratory of Professor Xi Jiang in the Division of Infectious Diseases, Cincinnati Children’s Hospital Medical Center, University of Cincinnati College of Medicine, Cincinnati, Ohio USA.
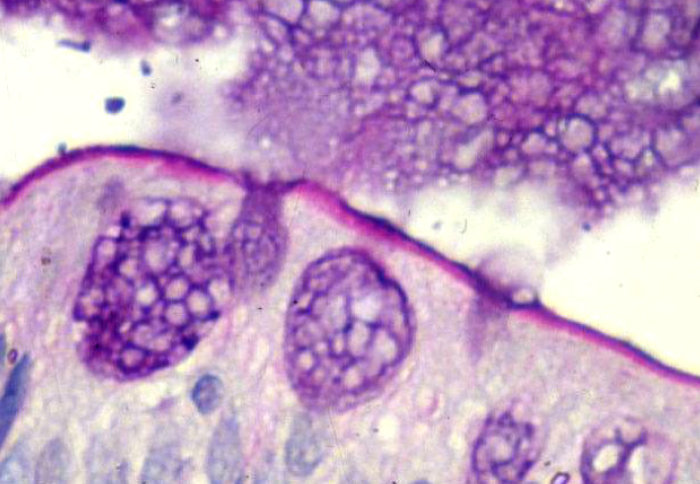
Main article image credit: Dr Alan Phillips.
The image depicts a small intestinal epithelial cell. PAS/Alcian Blue staining shows mucus (dark blue) in goblet cells and discharged into the Intestinal lumen and brush border staining (magenta) with mucus layer on the surface.
Article supporters
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Ms Genevieve Timmins
Academic Services