Research in infectious diseases from across the Faculty of Natural Sciences

The FoNS seminar took place on 28 February and this term's theme was Infectious Diseases.
The termly FoNS seminar brings together our community - students, academic staff, support staff and visitors - for a series of research talks followed by an informal drinks and canapé reception. It provides a great opportunity for early career researchers to present their work to a varied audience and for academics from across our departments to share their current research on a topical theme. All welcome!
The most recent seminar took place on 28 February, when around one hundred staff and students gathered together to hear presentations by seven researches from across the Faculty of Natural Sciences about their varied and cutting-edge work in the field of infectious diseases.
Oliver Bardell-Cox: Tackling bacterial resistance with the reverse antibiotics – the Nybomycins
First up was Oliver Bardell-Cox from the Department of Chemistry and the Institute of Clinical Sciences:
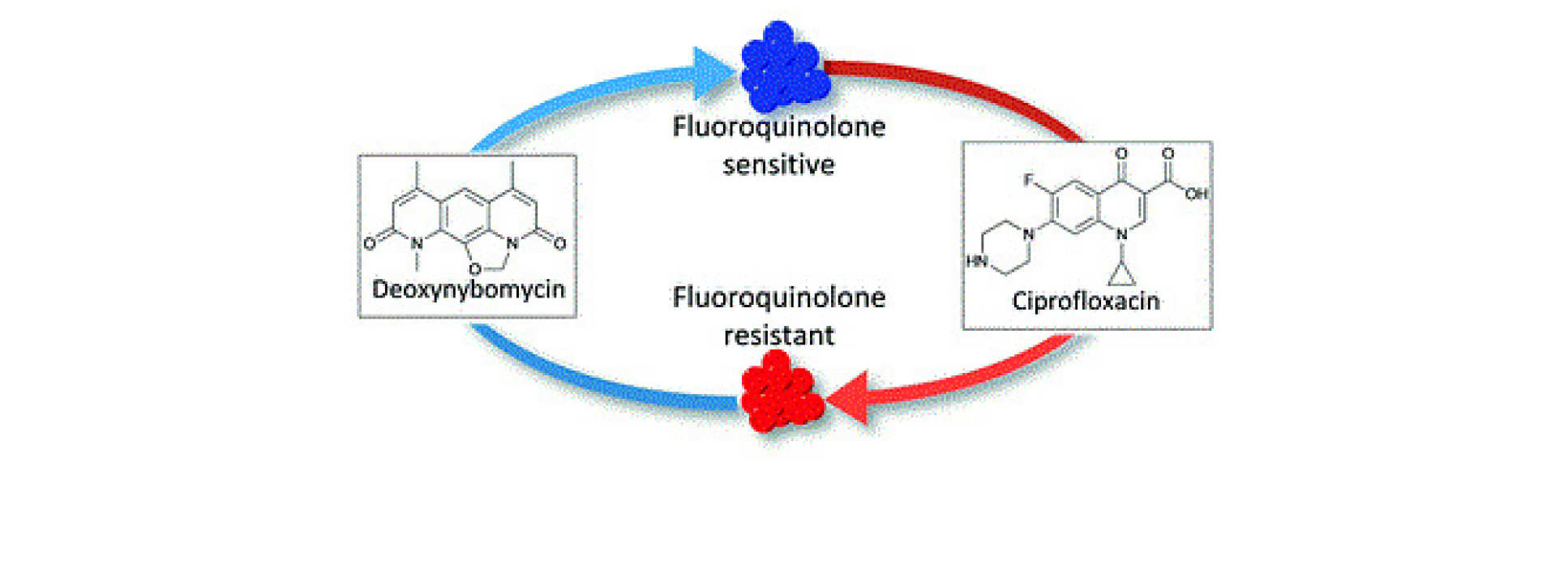
Abstact: The Nybomycins are a family of natural products that possess intriguing properties. They have the potential to extend the clinical efficacy of Fluoroquinolones, one of the most proscribed classes of antibiotics through a 'reverse antibiotic' approach. However, very little is known about the structure-activity relationship that underpins this approach. This talk discusses our efforts towards understanding what underpins this approach and how the Nybomycins family can be progressed into a therapeutic option.
Catherine Ducker: Investigating immune cell responses to helminth infection
Catherine Ducker from the Department of Life Sciences was our second speaker:
Abstract: Helminth parasites infect both humans and livestock, causing malnutrition and subsequent socio-economic effects. In order to establish chronic infection, helminths suppress the anti-parasite immune response. This response is co-ordinated by T-cells, a type of lymphocyte. However, the underlying mechanism is not fully known. Here we investigated T-cell responses to the natural murine gastrointestinal helminth, Heligmosomoides polygyrus (H. polygyrus). We analysed T-cell responses during infection using a fluorescent-timer reporter system, which allows analysis of temporal dynamics of gene activities in individual cells. We identify that during early infection, a proportion of parasite-reactive T-cells start to use the Foxp3 gene, which is specific to immune-suppressive T-cells (regulatory T-cells). Gene expression analysis reveals that T-cells with parasite-induced Foxp3 expression have characteristics of both regulatory T-cells and anti-parasite T-cells. We therefore propose the model that H. polygyrus manipulate anti-parasite T-cells to convert into immune-suppressive T-cells, specifically preventing the anti-parasite T-cell response. Consequently, H. polygyrus maintain chronic infection.
Dr Anna Barnard: Mimicking Protein-Protein Interactions in Persistent Bacteria
Dr Anna Barnard (Department of Chemistry) was next up, speaking about some of her group's recent research:
Abstract: Throughout biology, protein functions are critically mediated through interactions with other proteins.1 Some of the most interesting interactions are critical for pathogen survival, making PPIs promising targets for antimicrobial development.2 A major class of currently untargeted PPIs are toxin-antitoxin interactions which underlie bacterial persistence, a frequent cause of antibiotic treatment failure and relapsing infections.3 Unlike antibiotic resistance, whereby a genetic mutation allows bacteria to grow in the presence of antibiotics, persistence denotes the ability of bacteria to form persisters, a dormant non-growing phenotype tolerant to antibiotics.5 Antibiotic tolerance and ability to reinitiate growth after antibiotic removal are increasingly recognised as prevalent causes of bacterial infection relapses.3 Bacterial toxin-antitoxin (TA) operons are a ubiquitous feature of pathogenic bacteria and, when expressed, form a harmless toxin-antitoxin PPI. Under environmental or antibiotic stress the antitoxin is rapidly degraded, releasing the toxin which then inactivates bacterial growth, leading to the formation of persisters.4 The toxin Doc is a Fic domain kinase that induces dormancy by targeting protein translation through phosphorylation and inactivation of EF-Tu. Its action is inhibited through interaction with the α-helical C-terminal domain of antitoxin Phd.5
This presentation describes our efforts towards the first full characterisation of the Doc/Phd PPI in Salmonella and the development of a robust expression method for Doc toxins, applicable across all bacteria species. Our first-in-class peptide based Doc inhibitors that mimic the native antitoxin, Phd will also be discussed. This work clearly demonstrates the therapeutic potential of targeting toxin proteins, enabling clearance of persistent infections by restoring vulnerability to antibiotic treatment.
References:
1) M.P.H. Stumpf, T.Thorne, E. de Silva, R. Stewart, H.J. An, M. Lappe, C. Wiuf, Proc. Natl. Acad. Sci. U. S. A. 2008, 105, 6959.
2) L.-G. Milroy, T.N. Grossmann, S. Hennig, L. Brunsveld, C. Ottmann, Chem. Rev. 2014, 114, 4695.
3) A. Harms, E. Maisonneuve, K. Gerdes, Science 2016, 354, 1390.
4) R.A. Fisher, B. Gollen, S. Helaine, Nat. Rev. Microbiol. 2017, 15, 453.
5) A. Garcia-Pino, S. Balasubramanian, L. Wyns, E. Gazit, H. De Greve, R.D. Magnuson, D. Charlier, N.A.J. van Nuland, R. Loris, Cell 2010, 142, 101
Dr Gerald Larrouy-Maumus: Use of mass spectrometry to tackle AMR by improving diagnostic solutions and deciphering bacterial drug tolerance
Just before the break we heard from Dr Gerald Larrouy- Maumus (Department of Life Sciences):
Abstract: Antimicrobial resistance (AMR) is one of the biggest public health concerns of our time. There is a pressing need to decipher the mechanism leading to the raise of drug resistance in order to develop “smart antimicrobials” and also a need to determine quicker the antimicrobials susceptibility of bacterial pathogens in order to avoid the spread of the resistance. In our lab, by using mass spectrometry approaches, we have been able address those by showing that the small signalling molecule, cAMP, is involved in the process of antimicrobials tolerance in mycobacteria, which pathway represents a new and attractive route for antimicrobials development. As well, we have been able to develop a rapid and affordable assay to detect bacteria which are resistant to colistin, one of the last resort antibiotics, which technology can revolutionise drug susceptibility testing in clinical laboratories.
Dr Oliver Ratmann: Quantifying HIV transmission flow from cross-sectional viral phylogenetic deep sequence data: a population-based study in Rakai, Uganda
Dr Oliver Ratmann from the Department of Mathematics then presented his recent research examining HIV 'hotspots':
Abstract: To prevent new infections with human immunodeficiency virus type 1 (HIV-1) in sub-Saharan Africa, UNAIDS recommends targeting interventions to populations that are at high risk of acquiring and passing on the virus. However, it is often unclear who and where these “source” populations are. Viral deep-sequence data have recently been established as a method for inferring the direction of transmission at an accuracy sufficient for population-level analyses. We present a novel semi-parametric Bayesian modelling framework of viral sequence data to quantify HIV transmission flows and detect source and sink population groups that directly incorporates this information. Our model uses a high-dimensional set of Poisson regression equations, adjusting for sampling heterogeneity known from cross-sectional surveillance to avoid selection bias in the flow estimates. We illustrate the methodology on population-based sequence data obtained from infected individuals participating in the Rakai Community Cohort Study in south-eastern Uganda, indicating that the method provides new opportunities for identifying the drivers of HIV spread and designing effective public health interventions.
Dr Abigail Clements: Dissecting the unique virulence mechanisms of the emerging human pathogen Shigella sonnei
Dr Abigal Clements from the Department of Life Sciences then presented her research into Shigellosis:
Abstract: Shigellosis is an infection causing moderate to severe diarrhea, primarily in children under 5 years of age. There are estimated to be over 150,000 deaths attributed to Shigella worldwide. Shigellosis is mainly caused by two related bacterial species, Shigella flexneri and Shigella sonnei. S. sonnei is predominantly responsible for dysentery in developed countries, and is replacing S. flexneri in areas undergoing economic development and improvements in water quality. The vast majority of research into the molecular mechanisms of infection has been conducted using S. flexneri. We have been working to identify and characterise the differences between S. flexneri and S. sonnei infections that may explain the different epidemiology of these pathogens. Unique virulence determinants and metabolic characteristics of S. sonnei suggest a substantially different infection mechanism which could influence vaccine design, antibiotic development and public health initiatives.
Professor Ed Tate: Tackling drug resistance in viruses, parasites and bacteria using chemical biology
Professor Ed Tate from the Department of Chemistry, was the final speaker. His group works on chemistry applied to viruses, bacteria and parasites which cause infectious diseases, aiming to understand how they manipulate their human host, and to use this knowledge to find new ways to break drug resistance.
Abstract: My group develops new tools and approaches in chemical biology, drug discovery and proteomics, and we apply them to challenges in biomedicine. I will present some of our research on infectious diseases caused by viruses (common cold, polio), parasites (malaria) and bacteria, where we have worked with colleagues in FoNS and in Medicine to overcome drug resistance, and to identify and exploit 'irresistible' host mechanisms which can block infection.
Join us next time and stay in touch!
Join us for the next FoNS seminar on the theme of space - happening summer 2020 - further details will be posted on the FoNS research pages shortly.
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Claudia Cannon
The Grantham Institute for Climate Change