New insights into the genetic defects that lead to heart failure
by Ellyw Evans

Individual genetic defects lead to heart failure in different ways, highlighting potential biomarkers for targeted treatment.
The molecular and cellular mechanisms that lead to heart failure in people with cardiomyopathy – disease of the heart muscle – are determined by the specific gene variant each patient carries.
These are the findings of a study published today in the journal Science and based on the first comprehensive single-cell analysis of cardiac cells from healthy and failing hearts.
This study conducted by an international consortium of 53 scientists, including researchers from Imperial’s National Heart and Lung Institute (NHLI), shows that cell type compositions and gene activation profiles change according to the genetic variants.
The investigators say the findings can inform the design of targeted therapies that take into account each patient’s underlying gene defect responsible for their particular form of cardiomyopathy.
Dr Michela Noseda, co-senior author and a lecturer in Cardiac Molecular Pathology at NHLI, said: "Our study provides a paradigm shift demonstrating that not all heart failure is the same, suggesting patients might benefit from specific treatments. To move towards precision and interceptive medicine we now need to see what changes we discover in earlier stages of the disease."
The team studied 880,000 single heart cells
The interdisciplinary team from six countries examined the genes activated in about 880,000 single cells from 61 failing hearts and 18 healthy donor hearts as reference. Some diseased hearts were provided by the Cardiovascular Biomedical Research Unit Biobank, Royal Brompton and Harefield Hospitals.
Each cardiac cell type and the numerous subtypes from the sample hearts were analysed one by one using single-cell sequencing methods. No lab on its own could deal with the enormous amount of data generated, but close collaboration among specialists from different disciplines made it possible to assemble a coherent picture out of each individual piece of the puzzle.
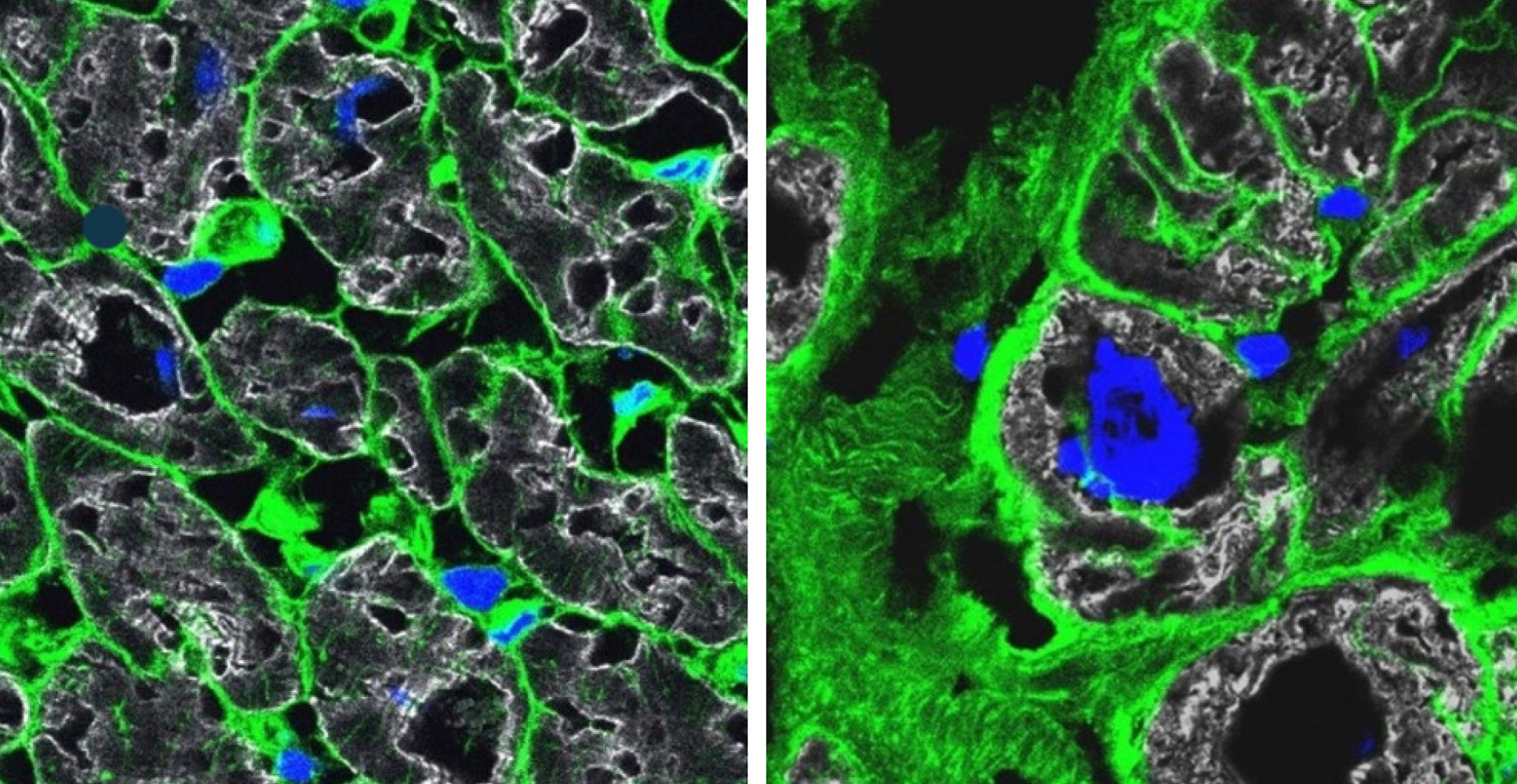
This study is also part of the efforts of the international Human Cell Atlas (HCA) consortium, which is aiming to map every cell type in the human body as a basis for both understanding human health and for diagnosing, monitoring, and treating disease.
Cardiomyopathy is not a uniform disease
The study focused on dilated cardiomyopathy (DCM), the most common form of heart failure that leads to heart transplantations. It involves an expansion of the walls of the heart chamber, particularly in the left ventricle, the heart’s main pumping chamber. The muscles of the heart become weakened, compromising its ability to contract and pump blood, which ultimately leads to heart failure.
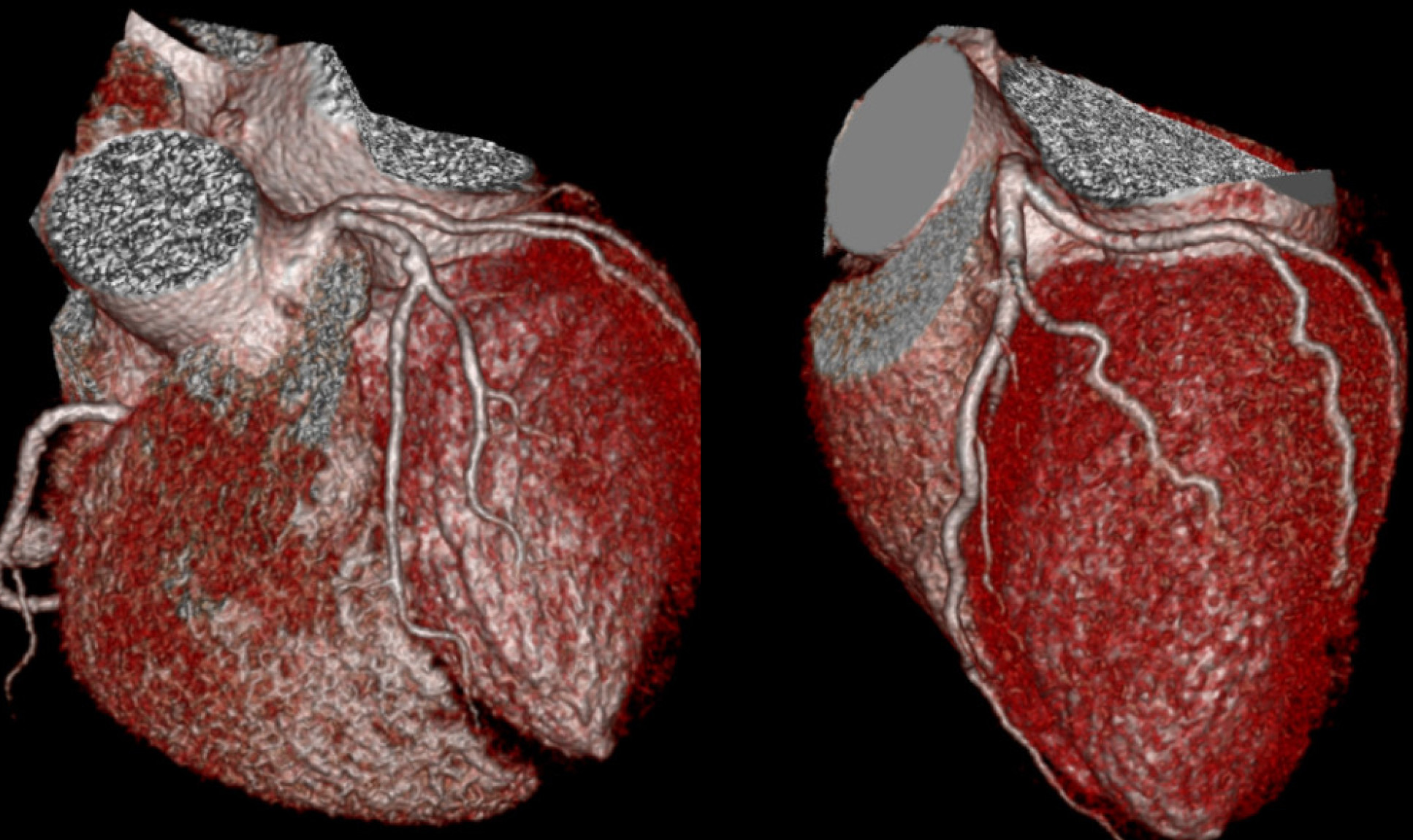
The consortium studied tissues from patients with different genetic mutations that commonly lead to cardiomyopathies. These mutations occurred in proteins with different functions in the heart and the analyses indicate that these triggered different responses.
Co-senior author Professor Norbert Hübner from the Max Delbrück Centre for Molecular Medicine Berlin, said: “We investigated pathogenic gene variants in heart tissue at the single-cell level, which allowed us to map precisely how specific pathogenic variants drive cardiac dysfunction. To our knowledge, this is the first such analysis conducted in cardiac tissue, and we hope this approach can be used to study other types of genetic heart diseases.”
Biomarkers for targeted therapies
According to the researchers, the ultimate goal is to develop individualised therapies for heart disease because genotype-specific treatment could be more effective and with fewer side effects. The consortium has made all of its results available to the scientific community online. The authors hope that this resource propels studies by other groups to define new treatments that prevent heart failure, which today is an incurable disease.
All data from the study is available online.
‘Pathogenic variants damage cell compositions and single cell transcription in cardiomyopathies’ by Daniel Reichart et al is published in Science.
This story was adapted from a press release by MDC Berlin.
Main image credit: Dr Eleonora Adami, MDC.
Article supporters
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Ellyw Evans
Faculty of Medicine Centre