Copying nature's lock-and-key system could improve rapid medical diagnostics
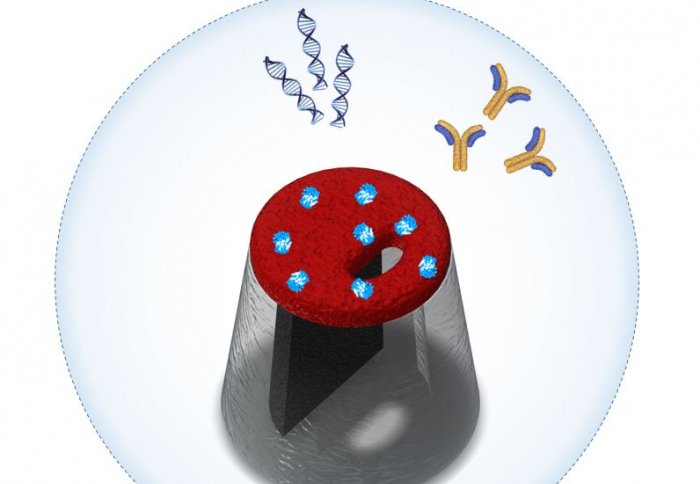
Researchers have designed a system that rapidly recognises the specific biological molecules that can indicate disease.
The team from Imperial College London have developed a nanoscale sensor that can selectively detect protein molecules at the single-molecule level, which could help in early stage clinical diagnosis.
The complete system combines concentration, tuneable speed and selectivity, which will be clinically relevant in the search for rare proteins such as specific kinds of antibodies and DNA molecules.
– Professor Yuri Korchev
When analysing body fluid samples for signals of a disease, scientists are often looking for very rare molecules within a complex mixture. In order to find such ‘needles in a haystack’, scientists often use methods that detect single molecules at a time.
One promising technology is nanopore sensing, where individual molecules are passed through a very small nanometer-sized hole. This process results in each molecule producing its own unique signature, without the need for lengthy sample preparation or chemical modification.
However, different molecules of the same size can produce very similar signals, making it hard to uniquely identify the target molecule.
To solve this problem, a team led by Imperial College London have developed a system based on a nanopore and a nanoscale transistor, which can recognise target molecules in a similar way to biological receptors. The details of their new system are published today in Nature Communications.
Lock and key
Receptors recognise molecules with particular shapes and bind to them in a lock-and-key mechanism. In this study, the nanoscale transistor was made from a polymer material that could be imprinted with a binding site – ‘the lock’. This allows the system to detect the only matching ‘key’ – a specific target molecule.
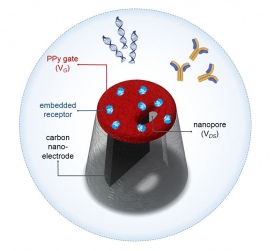 To test that the system works as expected, the team used it to detect the antibody that binds to insulin, a mechanism important in the diagnosis of diabetes. However, the team say that the system design can also be readily applied in the detection to a much broader range of biological molecules.
To test that the system works as expected, the team used it to detect the antibody that binds to insulin, a mechanism important in the diagnosis of diabetes. However, the team say that the system design can also be readily applied in the detection to a much broader range of biological molecules.
Co-author of the study Professor Joshua Edel, from the Department of Chemistry at Imperial, said: “We have shown that we can imprint a polymer at the entrance to the nanopore with the shape of the natural ‘lock’ to the ‘key’ molecule we are looking for, mimicking biological receptors.”
Opening and closing the gate
The researchers also added another feature to the new system to solve another problem in nanopore sensing: if molecules pass too quickly through the nanopore, they may not get detected.
They added an electrode attached to the polymer coating of the pore, forming a nanoscale transistor, to which a voltage could be applied. This makes the pore act like a gate – the applied voltage can ‘open’ or ‘close’ the gate, controlling the transport of molecules through the pore.
Dr Aleksandar Ivanov, from the Department of Chemistry at Imperial, said: “We now have a truly tuneable biosensor. By adding new complexity into the system we can control the transport of molecules and can have longer time to study a specific molecule.”
Professor Yuri Korchev, from the Department of Medicine at Imperial, added: “The complete system combines concentration, tuneable speed and selectivity, which will be clinically relevant in the search for rare proteins such as specific kinds of antibodies and DNA molecules.”
-
‘Nanopore Extended Field Effect Transistor for Selective Single Molecule Biosensing’ by Ren Ren, Yanjun Zhang, BinoyPaulose Nadappuram, Bernice Akpinar, David Klenerman, Aleksandar P. Ivanov, Joshua B. Edel and Yuri Korchev is published in Nature Communications.
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Hayley Dunning
Communications Division