Divert to blog, google scholar and OA
Topics
- Recent Highlights
- Functional molecular systems
- Coarse-grained modelling of DNA
- Thermodynamics of small systems
- Simulation tools and algorithms
- Biomolecular Engineering
Jan 2026 - Thermodynamic limits in far-from-equilibrium molecular templating networks published in Newton
Cells assemble complex molecules such as proteins using template polymers with information-carrying sequences. These sequences are used to direct the ordered assembly of a set of molecular building blocks to form exactly the right product, such as a specific protein, on demand. A crucial feature of this process is that the templates act catalytically, since they accelerate the assembly of a specific product without being consumed in the process. A single template can then be used to direct the assembly of many copies of a product. It has long been suspected that this transfer of information from catalytic templates to products must be associated with a minimal energy cost, in the same way that computation has lower bounds on energy input. To explore this hypothesis further, this work considers arbitrarily complex templating networks involving many assembly and disassembly processes. We show that the accuracy with which a specific set of products can be maintained is, indeed, constrained by energetic properties of the underlying network. Surprisingly, the operation of the most efficient templating networks is completely different from the behavior observed in nature, where molecules such as proteins are overwhelmingly assembled by templates and degraded in a distinct, template-free fashion. Instead, the most efficient approach, which both achieves the highest accuracy and has a negligible energetic cost, is to disassemble products via a reversal of the pathway by which they were formed. This observation raises interesting questions about why nature does not operate in this way and whether synthetic templating systems can be built that exploit this approach.
June 2025 - Information propagation through enzyme-free catalytic templating of DNA dimerization with weak product inhibition published in Nature Chemistry
Information propagation by sequence-specific, template-catalysed molecular assembly is a key process facilitating life’s biochemical complexity, yielding thousands of sequence-defined proteins from only 20 distinct building blocks. However, exploitation of catalytic templating is rare in non-biological contexts, particularly in enzyme-free environments, where even the template-catalysed formation of dimers is challenging. Typically, product inhibition—the tendency of products to bind to templates more strongly than individual monomers—prevents catalytic turnover. Here we present a rationally designed enzyme-free system in which a DNA template catalyses, with weak product inhibition, the production of sequence-specific DNA dimers. We demonstrate selective templating of nine different dimers with high specificity and catalytic turnover, then we show that the products can participate in downstream reactions, and finally that the dimerization can be coupled to covalent bond formation. Most importantly, our mechanism demonstrates a design principle for constructing synthetic molecular templating systems, a first step towards applying this powerful motif in non-biological contexts to construct many complex molecules and materials from a small number of building blocks.
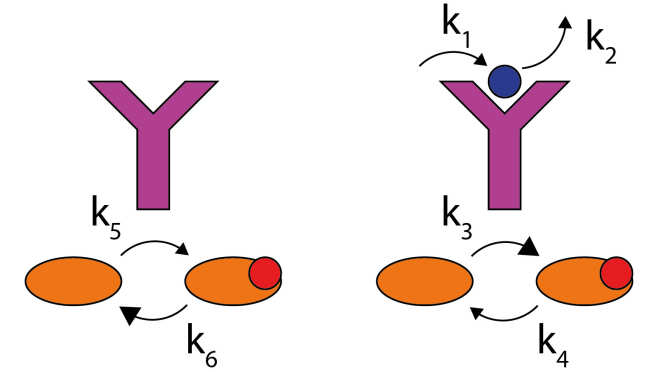
Biochemical networks within cells achieve remarkable functionality, including sensing, signalling, information-processing and replication. We aim to understand the fundamental physical principles that set the scope of this behaviour, allowing development of engineering principles for artificial analogs of these systems.
Since these systems of interest typically involve small numbers of molecules, randomness plays an important role. This work often touches on the deep connections between two bodies of work that deal explicitly with randomness: information theory, which describes the role of randomness in communication, and statistical mechanics, through which randomness is related to thermodynamics.
Recently, we've started to translate our basic understanding into the engineering of synthetic, nucleic acid-based analogs of these natural systems within our own lab here at Imperial. To get a taste, check out Javi's talk at FNANO2020, or a simplified discussion of his overall project.
Relevant Publications
- Plesa T, Dack A, Ouldridge TE, 2023 "Integral feedback in synthetic biology: Negative-equilibrium catastrophe", J. Math. Chem.
- Qureshi, B., Juritz, J., Poulton, J. M., Beersing-Vasquez, A., & Ouldridge, T. E. (2023). A universal method for analyzing copolymer growth. The Journal of Chemical Physics.
- JM Lawrence et al., 2022, Synthetic biology and bioelectrochemical tools for electrogenetic system engineering, Science Advances.
- Catala AC, Ouldridge TE, Stan GBV, Bae W, 2022, "Building an RNA-Based Toggle Switch Using Inhibitory RNA Aptamers", ACS Synthetic Biology.
- Juritz J, Poulton JM, Ouldridge TE, 2021, "Minimal mechanism for cyclic templating of length-controlled copolymers under isothermal conditions", The Journal of Chemical Physics.
- Bae W, Stan GBV, Ouldridge TE, 2021, "In situ generation of RNA complexes for synthetic molecular strand displacement circuits in autonomous systems", Nano Lett.
- Cabello-Garcia J, Bae W, Stan GBV, Ouldridge TE, 2021, "Handhold-mediated strand displacement: a nucleic acid-based mechanism for generating far-from-equilibrium assemblies through templated reactions", ACS Nano.
- Poulton JM, Ouldridge TE, 2021, "Edge-effects dominate copying thermodynamics for finite-length molecular oligomers", New J. Phys.
- Lankinen A, Mullor Ruiz I, Ouldridge TE, 2020, "Implementing Non-Equilibrium Networks with Active Circuits of Duplex Catalysts", 26th International Conference on DNA Computing and Molecular Programming (DNA 26).
- Plesa T, Stan G-BV, Ouldridge TE, Bae W, 2021, "Quasi-robust control of biochemical reaction networks via stochastic morphing", J. R. Soc. Interface.
- Deshpande A, Ouldridge TE, 2020, "Optimizing enzymatic catalysts for rapid turnover of substrates with low enzyme sequestration". Biological Cybernetics.
- Brittain RA, Jones NS, Ouldridge TE, 2019, Biochemical Szilard engines for memory-limited inference, New J. Phys.
- Poulton J, ten Wolde PR and Ouldridge TE, 2019, Non-equilibrium correlations in minimal dynamical models of polymer copying, PNAS.
- Ouldridge TE, Brittain RA, ten Wolde PR, 2018, The power of being explicit: demystifying work, heat, and free energy in the physics of computation, in "The Energetics of Computing in Life and Machines", SFI Press.
- Deshpande A, Ouldridge TE, 2017, High rates of fuel consumption are not required by insulating motifs to suppress retroactivity in biochemical circuits, Engineering Biology.
- Poole W, Ortiz-Muñoz A, Behera A, Jones NS, Ouldridge TE, Winfree E, Gopalkrishnan M, 2017, Chemical Boltzmann Machines, In DNA Computing and Molecular Programming (DNA 2017).
- Brittain RA, Jones NS, Ouldridge TE, 2017, What we learn from the learning rate, J. Stat. Mech.
- Ouldridge TE, 2017, The importance of thermodynamics for molecular systems, and the importance of molecular systems for thermodynamics, Nat. Comput.
- Ouldridge TE, ten Wolde PR, 2017, Fundamental costs in the production and destruction of persistent polymer copies, Phys. Rev. Lett.
- Ouldridge TE, Govern CC, ten Wolde PR, 2017, Thermodynamics of computational copying in biochemical system, Phys. Rev. X.
- McGrath T, Jones NS, ten Wolde PR, Ouldridge TE, 2017, Biochemical machines for the interconversion of mutual information and work, Phys. Rev. Lett.
- ten Wolde PR, Becker NB, Ouldridge TE, Mugler A, 2015, Fundamental Limits to Cellular Sensing, J. Stat. Phys.
- Ouldridge TE, ten Wolde PR, 2014, The robustness of proofreading to crowding-induced pseudo-processivity in the MAPK pathway, Biophysical Journal.
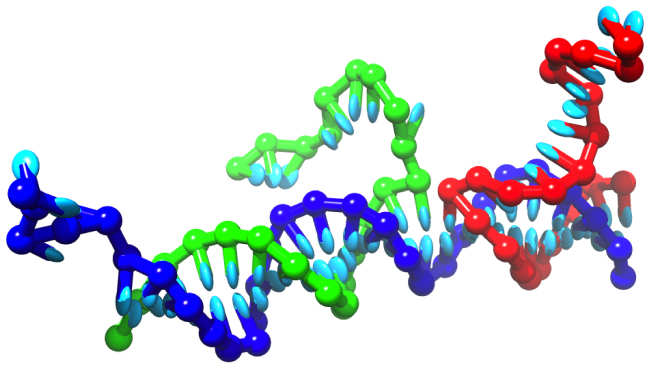
The elegant selectivity of Watson-Crick base-pairing makes DNA an extremely useful tool for the construction of nanoscale objects and machines. Stable structures and mechanical cycles can be programmed into a system of single strands by careful choice of the sequences of bases. I'm particularly interested in using nucleic acids to design artificial analogs of complex cellular systems, to enable careful exploration of the design principles and engineering possibilities.
Despite the experimental successes, there is no clear theoretical description of the processes involved. We have developed a nucleotide-level coarse grained model of DNA, oxDNA, which is detailed enough to capture the essential physics of assembly processes, yet simple enough to be applicable over long time scales. Code, user guides and examples for simulating the model can be downloaded from this site.
The oxDNA model was developed in the Doye / Louis groups in Oxford. It has since been applied in collaboration with the Turberfield group in Oxford, the Winfree group in Caltech and the Nir group at the Ben-Gurion University of Negev, as well as being used independently by other researchers.
Even with oxDNA, it is still not practical to simulate the formation of very large structures. A collaboration with the Turberfield and Kwiatkowska groups in Oxford has led to a less detailed model that can describe the formation of DNA origami structures.
Relevant Publications
- Sengar A, Ouldridge TE, Henrich O, Rovigatti L, Sulc P, 2021, "A Primer on the oxDNA Model of DNA: When to Use it, How to Simulate it and How to Interpret the Results", Frontiers in Molecular Biosciences.
- Irmisch P, Ouldridge TE, Seidel R, 2020, "Modelling DNA-strand displacement reactions in the presence of base-pair mismatches", J. Am. Chem. Soc.
- Haley NEC, Ouldridge TE, Mullor Ruiz I, Geraldini A, Louis AA, Bath J and Turberfield AA, 2020, Design of hidden thermodynamic driving for non-equilibrium systems via mismatch elimination during DNA strand displacement, Nat. Comms.
- Fonseca P, Romano F, Schreck JS, Ouldridge TE, Doye JPK and Louis AA, 2018, Multi-scale coarse-graining for the study of assembly pathways in DNA-brick self assembly, J. Chem. Phys.
- Khara DC, Schreck JS, Tomov TE, Berger Y, Ouldridge TE, Doye JPK and Nir E, 2018, DNA bipedal motor walking dynamics: an experimental and theoretical study of the dependency on step size, Nucl. Acids Res.
- Snodin BEK, Romano F, Rovigatti L, Ouldridge TE, Louis AA, and Doye JPK, 2016, Direct Simulation of the Self-Assembly of a Small DNA Origami, ACS Nano.
- Dunn KE, Dannenberg F, Ouldridge TE, Kwiatkowska M, Turberfield AJ, Bath J, 2015, Guiding the folding pathway of DNA origami, Nature.
- Dannenberg F, Dunn KE, Bath J, Kwiatkowska M, Turberfield AJ, Ouldridge TE, 2015, Modelling DNA Origami Self-Assembly at the Domain Level, J. Chem. Phys.
- Snodin BEK, Randisi F, Mosayebi M, Sulc P, Schreck JS, Romano F, Ouldridge TE, Tsukanov R, Nir E, Louis AA, Doye JPK, 2015, Introducing improved structural properties and salt dependence into a coarse-grained model of DNA, J. Chem. Phys.
- Schreck JS, Ouldridge TE, Romano F, Sulc P, Shaw L, Louis AA, Doye JPK, 2015, DNA hairpins primarily promote duplex melting rather than inhibiting hybridization, Nucleic Acids Research.
- Mosayebi M, Louis AA, Doye JPK, Ouldridge TE, 2015, Force-induced rupture of a DNA duplex, ACS Nano.
- Machinek RR, Ouldridge TE, Haley NE, Bath J, Turberfield AJ, 2014, Programmable energy landscapes for kinetic control of DNA strand displacement, Nature Communications.
- Doye JPK, Ouldridge TE, Louis AA, Romano F, Sulc P, Matek C, Snodin BEK, Rovigatti L, Schreck JS, Harrison RM, Smith WPJ, 2013, Coarse-graining DNA for simulations of DNA nanotechnology, Physical Chemistry Chemical Physics.
- Srinivas N, Ouldridge TE, Sulc P, Schaeffer JM, Yurke B, Louis AA, Doye JPK, Winfree E, 2013, On the biophysics and kinetics of toehold-mediated DNA strand displacement, Nucleic Acids Research.
- Ouldridge TE, Sulc P, Romano F, Doye JPK, Louis AA, 2013, DNA hybridization kinetics: Zippering, internal displacement and sequence dependence, Nucleic Acids Research.
- Ouldridge TE, Hoare RL, Louis AA, Doye JPK, Bath J, Turberfield AJ, 2013, Optimizing DNA nanotechnology through coarse-grained modeling: A two-footed DNA walker, ACS Nano.
- Sulc P, Romano F, Ouldridge TE, Rovigatti L, Doye JPK, Louis AA, 2012, Sequence-dependent thermodynamics of a coarse-grained DNA model, Journal of Chemical Physics.
- Ouldridge TE, Louis AA, Doye JPK, 2011, Structural, mechanical, and thermodynamic properties of a coarse-grained DNA model, Journal of Chemical Physics.
- Ouldridge TE, Louis AA, Doye JPK, 2010, DNA nanotweezers studied with a coarse-grained model of DNA, Physical Review Letters.
Thermodynamics, the science of heat and energy transfer, emerged as a field in the 19th century, motivated by the need to describe the engines that powered the industrial revolution. One of the challenges of modern science is to adapt and extend the theory to describe microscopic systems in which fluctuations play a key role. Biological and biologically-inspired systems are a key arena for these new ideas, due both to the need to understand natural molecular analogues of the engines and processes that we are familiar with at much larger length scales, and the possibility of developing artificial devices ourselves.
Not only does thermodynamics provide understanding of biological systems, but the study of real biophysical devices in turn provides us with a deeper understanding of the thermodynamic principles at play. In particular, the natural diffusive behaviour of biomolecules allows us to study complex systems that do not require external manipulation to function.
Relevant Publications
- Ouldridge TE and Wolpert DH, 2023. Thermodynamics of deterministic finite automata operating locally and periodically. New Journal of Physics.
-
Seet I, Ouldridge TE, Doye JPK, 2023, Simulation of reversible molecular mechanical logic gates and circuits, Physical Review E.
- Qureshi B, Juritz J, Poulton JM, Beersing-Vasquez A, Ouldridge, TE, 2023, A universal method for analyzing copolymer growth. The Journal of Chemical Physics.
- Juritz J, Poulton JM, Ouldridge TE, 2021, "Minimal mechanism for cyclic templating of length-controlled copolymers under isothermal conditions", The Journal of Chemical Physics.
- Poulton JM, Ouldridge TE, 2021, "Edge-effects dominate copying thermodynamics for finite-length molecular oligomers", New J. Phys.
- Ouldridge TE, 2020, A biochemical device to demystify a century-old thermodynamics puzzle from theoretical physics, Reseach Outreach.
- Brittain RA, Jones NS, Ouldridge TE, 2019, Biochemical Szilard engines for memory-limited inference, New. J. Phys.
- Ouldridge TW, Brittain RA, ten Wolde PR, 2019, The power of being explicit: demystifying work, heat, and free energy in the physics of computation, in "The Energetics of Computing in Life and Machines", SFI Press.
- Stopnitzky E, Still S, Ouldridge TE and Altenberg L, 2019, Physical Limitations of Work Extraction from Temporal Correlations, Phys. Rev. E.
- Poulton J, ten Wolde PR and Ouldridge TE, 2019, Non-equilibrium correlations in minimal dynamical models of polymer copying, PNAS.
- Deshpande A, Ouldridge TE, 2017, High rates of fuel consumption are not required by insulating motifs to suppress retroactivity in biochemical circuits, Engineering Biology: 1.
- Deshpande A, Gopalkrishnan M, Ouldridge TE, Jones NS, 2017, Designing the Optimal Bit: Balancing Energetic Cost, Speed and Reliability, Proc. Roy. Soc. A.
- Brittain RA, Jones NS, Ouldridge TE, 2017, What we learn from the learning rate, J. Stat. Mech.
- Ouldridge TE, 2017, The importance of thermodynamics for molecular systems, and the importance of molecular systems for thermodynamics, Nat. Comput.
- Ouldridge TE, ten Wolde PR, 2017, Fundamental costs in the production and destruction of persistent polymer copies, Phys. Rev. Lett.
- Ouldridge TE, Govern CC, ten Wolde PR, 2017, Thermodynamics of computational copying in biochemical system, Phys. Rev.
- McGrath T, Jones NS, ten Wolde PR, Ouldridge TE, 2017, Biochemical machines for the interconversion of mutual information and work, Phys. Rev. Lett.
Our work often involves systems that are too complex to be treated analytically. This means that simulations are a key tool in our research, and we are interested in simulation techniques and analysis tools for systems involving biomolecular reactions.
In this work we collaborate with the Doye / Louis groups in Oxford, the biochemical networks group of Pieter Rein ten Wolde in Amsterdam, Michael Tretyakov in Nottingham and Ruslan Davidchack in Leicester, and Oliver Henrich in Strathclyde.
Relevant Publications
- Qureshi B, Juritz J, Poulton JM, Beersing-Vasquez A, & Ouldridge TE, 2023, A universal method for analyzing copolymer growth. The Journal of Chemical Physics.
- Sengar A, Ouldridge TE, Henrich O, Rovigatti L, Sulc P, 2021, "A Primer on the oxDNA Model of DNA: When to Use it, How to Simulate it and How to Interpret the Results", Frontiers in Molecular Biosciences.
- Henrich O, Yair AGF, Curk T, Ouldridge TE, 2018, Coarse-grained simulation of DNA using LAMMPS, Eur. Phys. J. E.
- Davidchack RL, Ouldridge TE, Tretyakov MV, 2017, Geometric integrator for Langevin systems with quaternion-based rotational degrees of freedom and hydrodynamic interactions, J. Chem. Phys.
- Vijaykumar A, Ouldridge TE, ten Wolde PR, Bolhuis PG, 2017, Multiscale simulations of anisotropic particles combining Brownian Dynamics and Green's Function Reaction Dynamics, Journal of Chemical Physics.
- Davidchack RL, Ouldridge TE, Tretyakov MV, 2015, New Langevin and Gradient Thermostats for Rigid Body Dynamics, Journal of Chemical Physics.
- Ouldridge TE, 2012, Inferring bulk self-assembly properties from simulations of small systems with multiple constituent species and small systems in the grand canonical ensemble, Journal of Chemical Physics.
- Ouldridge TE, Louis AA, Doye JPK, 2010, Extracting bulk properties of self-assembling systems from small simulations, Journal of Physics: Condensed Matter.
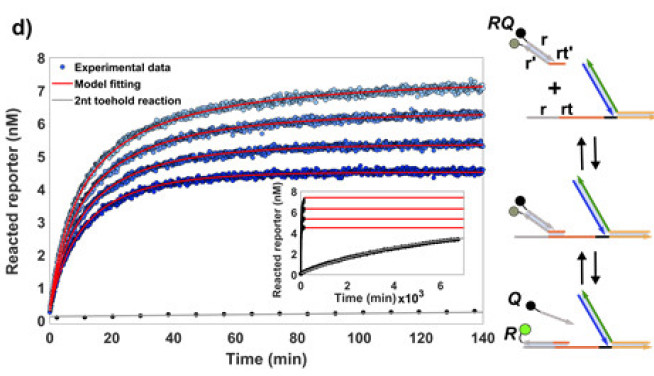
We've recently acquired our own (small) lab within the synthetic biology space at Imperial. We're using this lab to actually put our theoretical ideas into practice, engineering functional molecular systems from nucleic acids. These systems are both useful test-beds for our theory and engineering platforms for synthetic biology.
Our work currently focusses on engineering non-equilibrium information processing systems, analogs of the signalling and transcription/translation machinery in cells. To get a taste, check out Javi's talk at FNANO2020, or a simplified discussion of his overall project.
Below we list papers that present data obtained in our lab.
Relevant Publications
- Catala AC, Ouldridge TE, Stan GBV, Bae W, 2022, "Building an RNA-Based Toggle Switch Using Inhibitory RNA Aptamers", ACS Synthetic Biology,
- Bae W, Stan GBV, Ouldridge TE, 2021, "In situ generation of RNA complexes for synthetic molecular strand displacement circuits in autonomous systems", Nano Lett.
- Cabello-Garcia J, Bae W, Stan GBV, Ouldridge TE, 2021, "Handhold-mediated strand displacement: a nucleic acid-based mechanism for generating far-from-equilibrium assemblies through templated reactions", ACS Nano.
- Haley NEC, Ouldridge TE, Mullor Ruiz I, Geraldini A, Louis AA, Bath J and Turberfield AA, 2020, Design of hidden thermodynamic driving for non-equilibrium systems via mismatch elimination during DNA strand displacement, Nat. Comms.