Overview
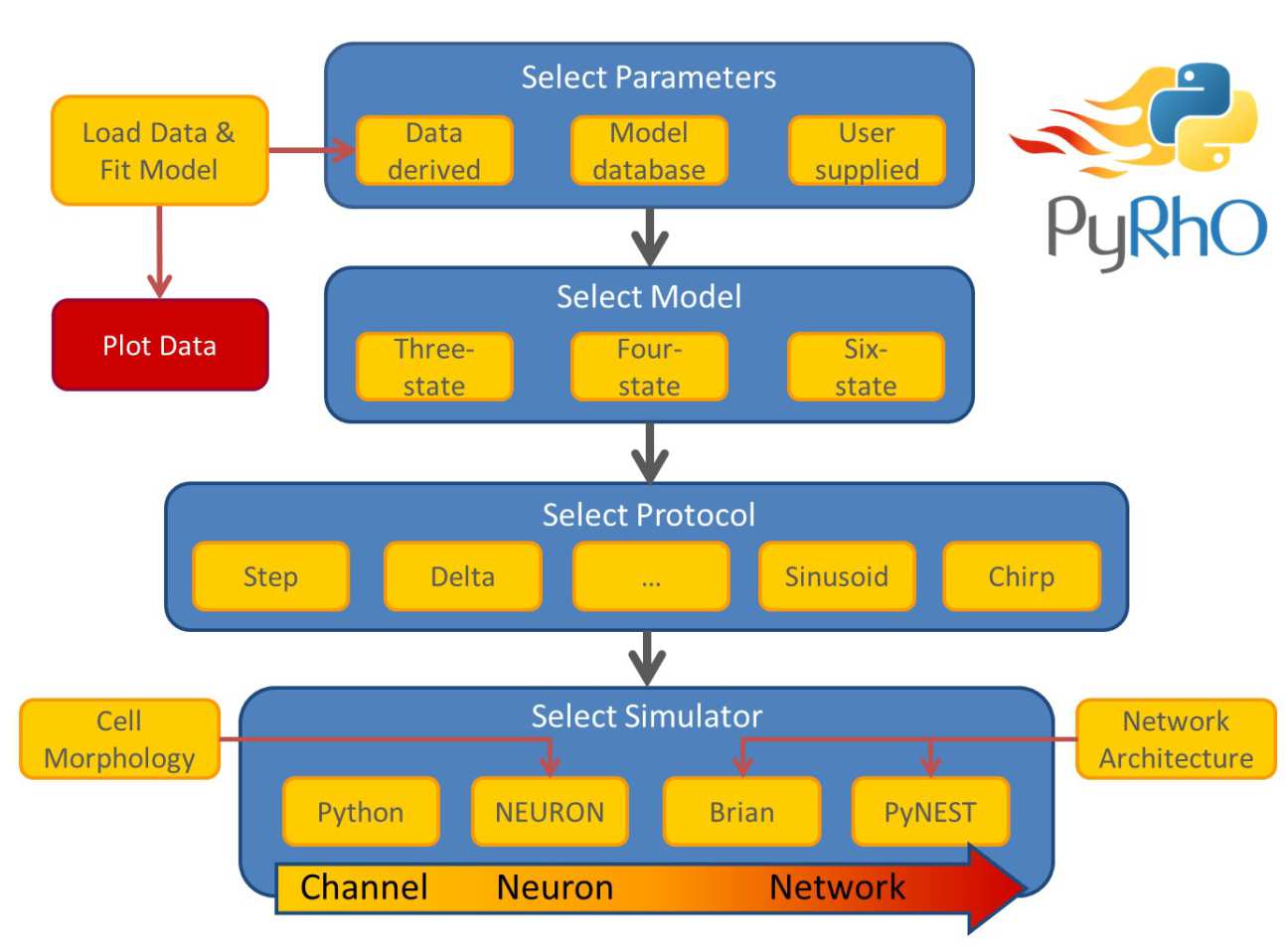
PyRhO is composed of several abstraction layers, including the Model layer, Protocol layer and Simulator layer (shown in the architectural schematic below). Choices in each layer are independent or one another, giving a large number of possible combinations to suit many needs for in silico experiments. Additionally, parameters may be fit to each type of kinetic model from experimental data or loaded from the default options of popular opsins.
For a more detailed introduction along with worked examples, please see our paper.
Abstraction Layers
| States | Transitions | Parameters | Pros | Cons |
|---|---|---|---|---|
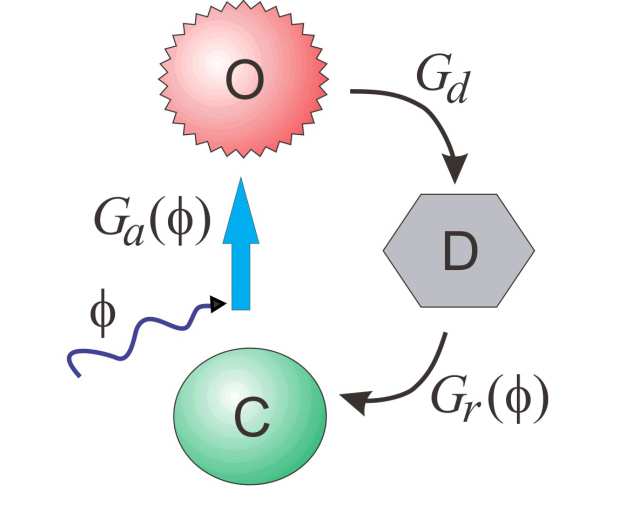
| 3 | 3 | 11 | Efficient analytic solution | Single exponential off-phase decay |
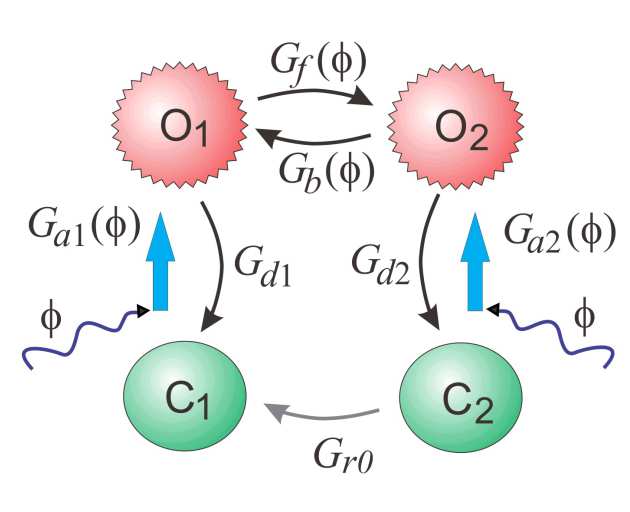
| 4 | 7 | 17 | Balance of detail and efficiency | Lacks short-pulse dynamics |
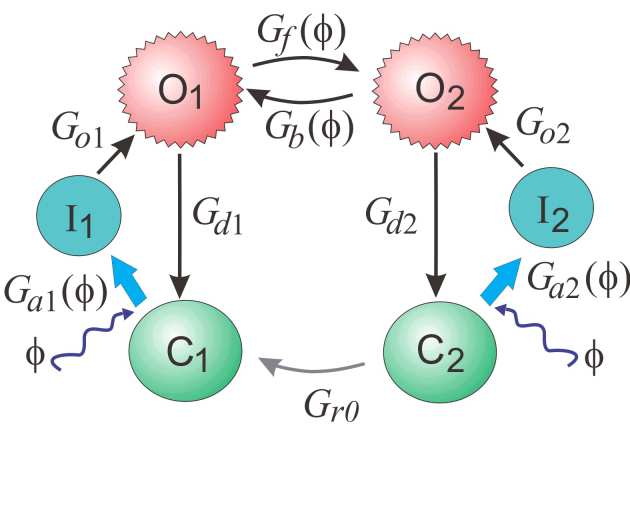
| 6 | 9 | 19 | Most detailed dynamics | Computationally expensive |
| Three-state model | Four-state model | Six-state model |
|---|---|---|
 |
 |
 |
PyRhO comes with several preconfigured and customisable simulation protocols for exploring the dynamics of the models. These include typical system analysis stimuli such as delta functions, step functions and sinusoids, as well as chirps and specialised protocols designed to probe particular features of the opsins including voltage-dependence (rectifier), opsin activation (shortPulse) and dark recovery (recovery).
| Name | Type | Purpose | Variations | Notes |
|---|---|---|---|---|
| step | Discrete | Standard pulse train | ||
| delta | Discrete | Quick perturbation | ||
| sinusoid | Continuous | Oscillating pulse | {sine, cosine} | |
| chirp | Continuous | Frequency analysis | {linear, exponential} | |
| ramp | Continuous | Linearly varying pulse | ||
| rectifier | Discrete | Assess voltage dependence | ||
| shortPulse | Discrete | Assess short pulse dynamics | ||
| recovery | Discrete | Assess dark recovery rate | ||
| custom | Continuous | Build custom stimulation function | User supplied |
Python
For opsin-level simulations, PyRhO solves the ordinary differential equations describing the opsin models with numerical integrators under a given set of experimental conditions. This assumes the opsin is essentially in something like a HEK cell where there are no confounding effects of other ion channels. The resultant photocurrents are then calculated and plotted.
NEURON
If more detailed morphological neuron models are desired, the topology, geometry and biophysics may be specified in a list of hoc files and passed to the NEURON simulator, along with the parameterised opsin model to be inserted into the cell's membrane. Additionally, the compartment name of where to record from should be specified (default: Vcomp='soma') along with the compartmental expression probability (default: expProb=1.0) and the initialisation membrane potential (default: v_init=-65) and whether to use a voltage clamp (default Vclamp=False). If desired, details of NEURON's numerical integrator may also be adjusted (defaults: CVode=False; dt=0.1) to balance the accuracy and runtime of the simulation.
Brian 2
For network level simulations, groups of spiking neurons may be transfected with a model opsin and simulated with the Brian 2 simulator. Brian is not currently integrated with PyRhO's GUI and so networks for simulation must be specified programmatically (although the opsin parameters may still be fit or loaded from the GUI). A worked example may be found in the demonstration notebook. For further details, examples and tutorials for using Brian, we refer you to the Brian 2 documentation.