What we do
Biostatistics
ECMC provides biostatistical support (e.g. sample size and power calculation, advice on the study design and statistical analysis plan) for early Phase I/II clinical trials or biomarker evaluation and target identification studies using clinical trial samples. Email queries to ecmcstatistics@imperial.ac.uk
Coordinate and run early phase trials
We have early phase, imaging and surgery trial coordinators who can help develop clinical trial protocols and run the trials.
Biobanking
We can help collect a range of samples from patients enrolled on clinical trials or undergoing standard care of treatment. Types of samples include surgical tissue, biopsy, whole blood, plasma, serum, urine, PBMCs and ascetic/cystic/pleural fluids.
Help researchers develop laboratory manuals so that samples are collected using standardised methods and using best practice.
BRC Circulating Tumour Biomarker
The BRC Circulating Biomarker Laboratory can analyse isolate and analyse single circulating tumour cells (CTCs) and cell-free circulating tumour DNA (ctDNA) from patient samples.
The laboratory is equipped with Cell Search; DEPArray NxT and has access to RainDrop Digital PCR system and Ion Chef/Ion S5 platforms.
Research Pathology Facility
We have histologist embedded in the Imperial Pathology Facility at Charing Cross Hospital who can provide:
- Cutting up
- Processing and embedding tissue
- Microtomy
- Haematoxylin
- Antibody optimisation
- Immunohistochemistry service
How to access ECMC support?
Who can apply?
Any Imperial College London and Imperial College Healthcare Trust staff involved in early translational cancer research.
Required Criteria for ECMC support.
The project work must meet ECMC remit which is early phase clinical trial/studies or biomarker evaluation and validation
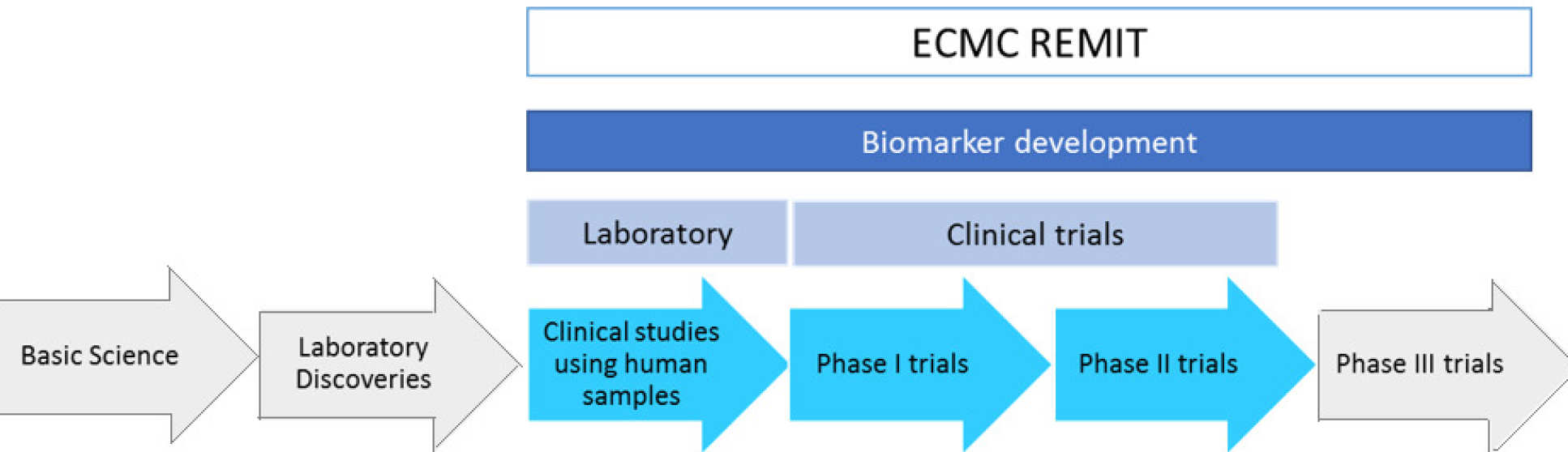
Research support application process
Who can apply? - Any cancer researchers at Imperial who is carrying out early phase clinical trials or biomarker evaluation.
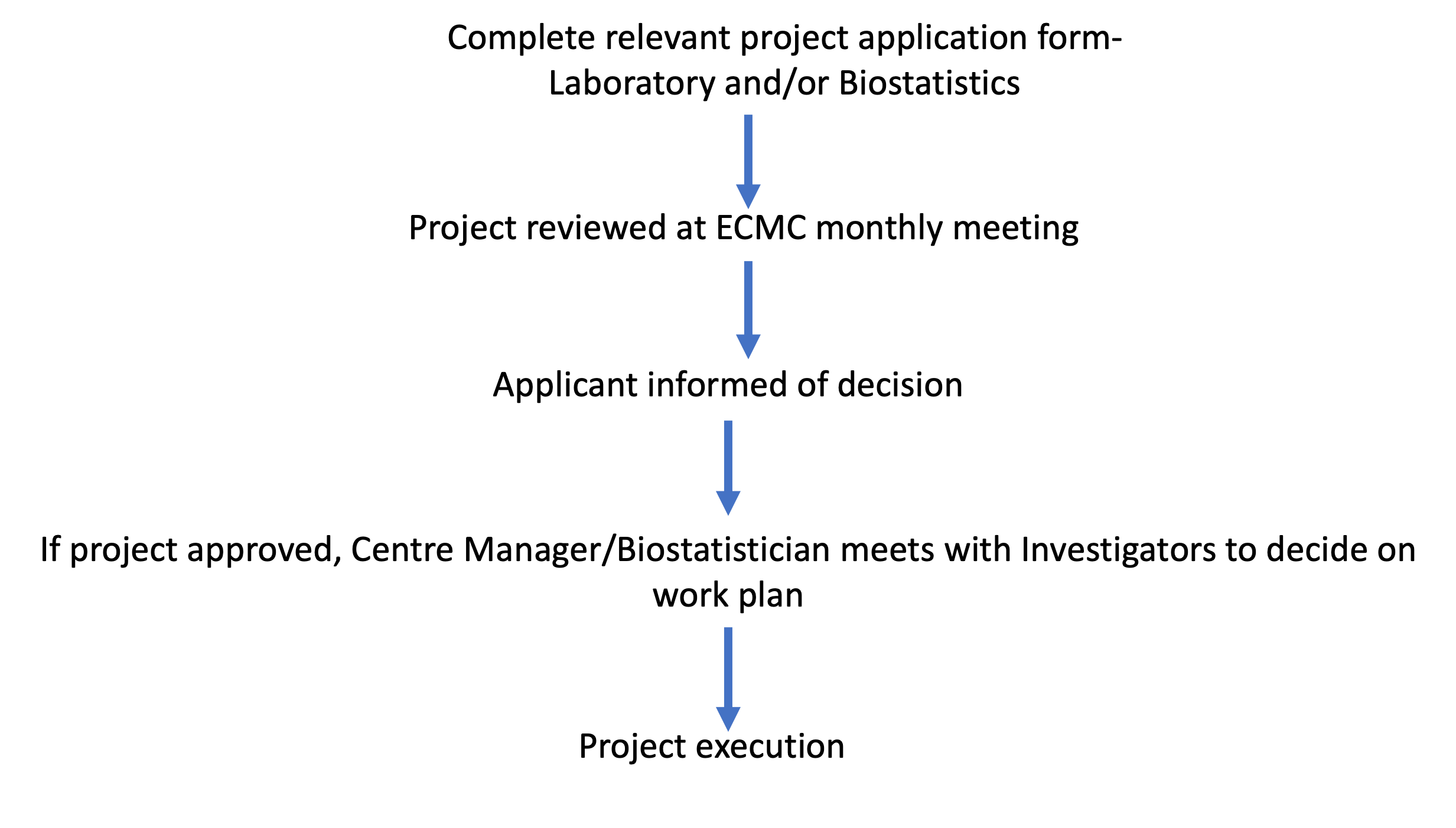
Download the application form (biostatistical support)
Download the application form (laboratory support)
ECMC Acknowledgement
Any work supported by Imperial ECMC must be acknowledged in study publications.
Suggested acknowledgement text- ‘This work is/was funded by grants from <funder1 and funder2>, and infrastructure support was provided by Imperial Experimental Cancer Medicine Centre, Cancer Research UK Imperial Centre, National Institute for Health Research (NIHR) Imperial Biomedical Research Centre (BRC) and Imperial College Healthcare NHS Trust Tissue Bank’
Patient access
Funding bodies expect public involvement in your research activities. By involving patients in your research, you can ensure that your research plans remain relevant to patient interest, that they are feasible and acceptable to potential research participants and that you communicate and disseminate your research in an accessible way to the general public.
The Imperial Involvement Group for Cancer can help in the following ways:
- help to co-produce a lay summary
- help to co-produce a Patient Information Sheet
- provide input into grant applications
- help you disseminate your research findings to the public is an accessible format
If you would like to discuss your needs in further detail, please contact Kelly Gleason.
Find a list of organisations and contacts who can support you with this.



