Tuberculosis genetic breakthrough may help tackle drug resistance
by Kate Wighton
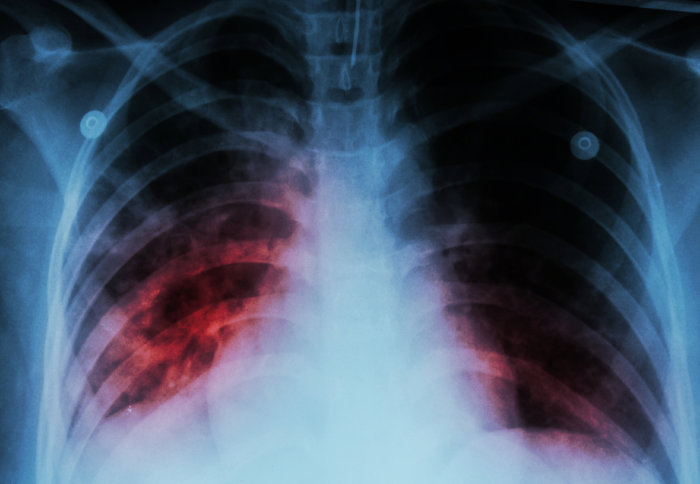
Imperial scientists have helped unravel the genetic code of tuberculosis (TB), enabling doctors to tailor treatments to each individual patient.
The study, published in the New England Journal of Medicine, may herald a quicker, more tailored treatment for the millions of people around the world living with tuberculosis.
The research suggests our understanding of TB’s genetic code is now so detailed we can predict which commonly used anti-TB drugs are best for treating a patient’s infection.
Developing more effective approaches to treating multi-drug-resistant TB is crucial for the thousands of people affected in the UK and millions worldwide Professor Chris Whitty Chief Scientific Adviser for the Department of Health and Social Care
This study, led by the international CRyPTIC consortium based at the University of Oxford and facilitated by the United Kingdom government’s 100,000 Genomes Project in partnership with Public Health England, is by far the largest of its kind, covering over 10,000 TB genomes from 16 equal partner countries around the globe.
The study, which also involved scientists from the University of Leeds and London School of Hygiene & Tropical Medicine, revealed a much greater accuracy in predicting the susceptibility of the bacterium to anti-TB drugs than had been expected.
Tuberculosis remains the world’s biggest infectious disease killer, claiming 1.7 million lives in 2016. The number of drug-resistant cases is rising, meaning new strategies and interventions are urgently needed if the World Health Organisation’s (WHO) target to end the global TB epidemic by 2035 is to be met.
One of the key interventions for achieving this target – and saving millions of lives - is getting the correct drugs to patients in a timely manner.
Since TB antibiotics were first introduced 70 years ago, tests to determine which antibiotics will best treat an individual patient have depended on growing the bacteria in a laboratory, a process that takes weeks or even months. These difficult and slow tests remain out of reach for most of the world’s TB patients, leaving many on the wrong combination of drugs and with a reduced chance of cure and survival.
It is estimated that only 22 per cent of an estimated 600,000 patients requiring treatment for multi-drug-resistant tuberculosis received diagnoses and were treated in 2016, which has facilitated the spread of multidrug-resistant strains.
No more guess work
The analysis conducted by the researchers, who looked at all major strains of TB, suggests that whole-genome sequencing can now characterise profiles of susceptibility to first-line anti-tuberculosis drugs, with a degree of accuracy sufficient for clinical use.
The lead investigator, Dr Tim Walker, Academic Clinical Lecturer in Microbiology and Infectious Diseases at the University of Oxford’s Nuffield Department of Medicine, said: “This study represents a paradigm shift away from a dependence on testing drugs against bacteria in culture and towards the genetic era.
“With ever-faster and more portable DNA sequencing technologies being developed, this advance means that we are now much closer to delivering tailored therapy to TB patients around the world whose treatments have so far been largely based on a ‘best guess’. Giving the correct drugs to more patients will improve cure rates and help stop the spread of drug-resistant strains,”
Francis Drobniewski, Professor of Global Health and Tuberculosis at Imperial’s Department of Medicine, and an author on the study added: “My team was delighted to participate in this study, bringing in genomic and microbiological data from the UK and East Europe - where the problem of multi-drug resistant tuberculosis has been particularly severe.
"When we published our population-based 1000 patient study in 2014 of Russian TB patients, we were not sure that we would be able to use whole genome sequencing (WGS) as a potential replacement for bacteriological-based drug resistance testing within just a few years."
Professor Drobniewski added: "At the moment we are still producing WGS from bacteriological cultures. The next stage will be to reliably perform WGS on material directly obtained from the patient, as close to the patient as possible”
Better treatments
Professor Chris Whitty, Chief Scientific Adviser for the Department of Health and Social Care, said: “Developing more effective approaches to treating multi-drug-resistant TB is crucial for the thousands of people affected in the UK and millions worldwide. This study is just one example of how the government is supporting research into how new technologies can help us tackle drug-resistant infections and thus preserve the effectiveness of current antibiotic treatments.”
The results of this study have already led to decisions in England, the Netherlands and by the Wadsworth Centre for Public Health, New York State, to reduce reliance on bacterial culture and deliver treatment according to DNA sequencing results alone.
This international research was supported in the UK by the Department of Health and Social Care through the National Institute for Health Research, Public Health England and the 100,000 Genomes Project. The research also received support from the Medical Research Council, the Wellcome Trust, and the Bill and Melinda Gates Foundation.
Article supporters
Article text (excluding photos or graphics) © Imperial College London.
Photos and graphics subject to third party copyright used with permission or © Imperial College London.
Reporter
Kate Wighton
Communications Division