Our research interests
- Our general interests
- Electron cryo-tomography
- Evolutionary analysis
- Microbiology
- Method Development
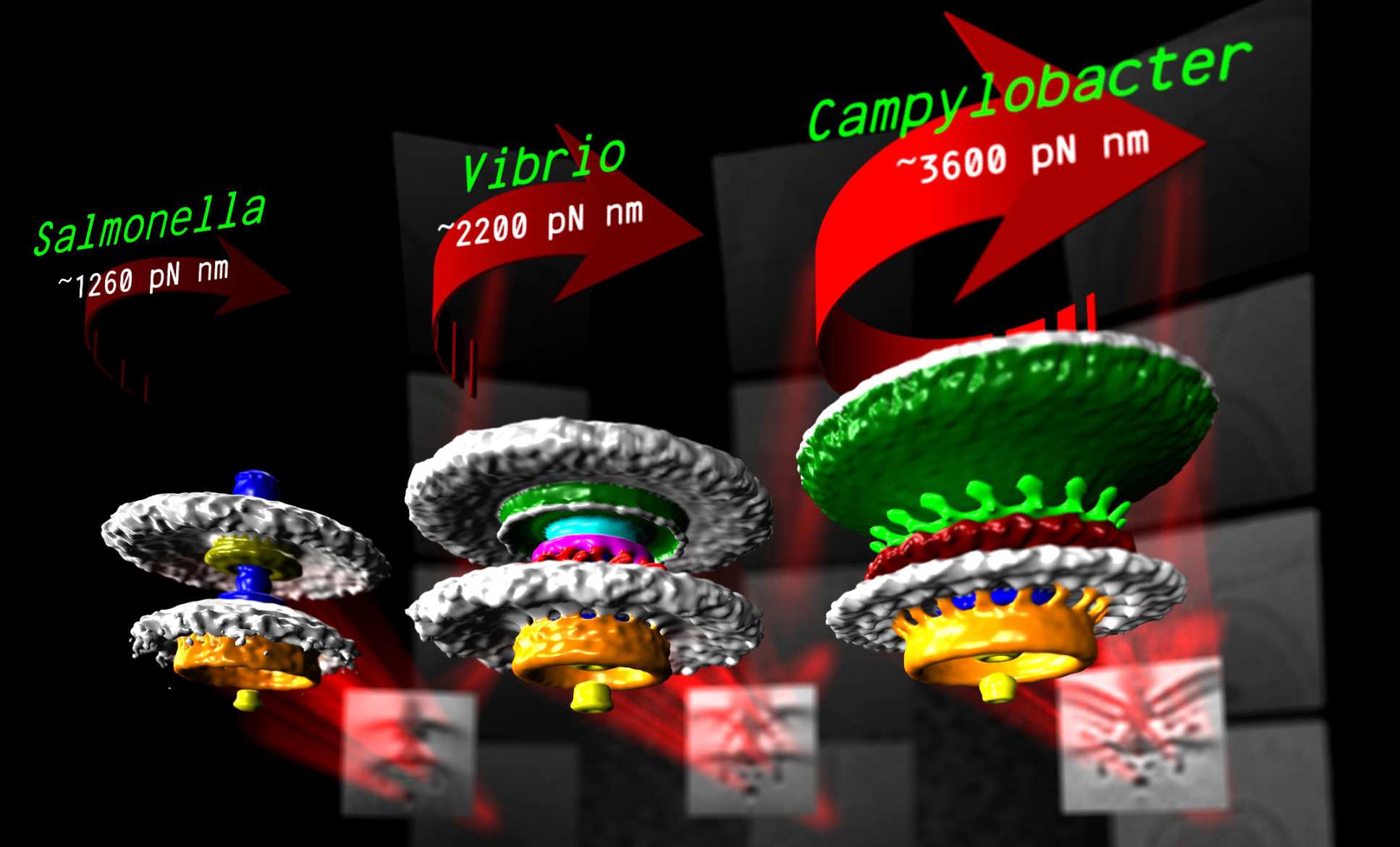

We are interested in how the molecular machines of the cell assembles, functions, and evolves. To tackle this problem we use electron cryo-tomography, a technique that enables us to visualize this machinery inside living cells -- to resolutions capable of discerning individual proteins. The technique involves flash-freezing the specimen then imaging it over a range of angles in an electron microscope. The resulting images can then be used to detemine the 3-D structure of the specimen in a manner directly analogous to CT or CAT scans. We interpret our imaging results in the light of molecular phylogenetics to understand the evolutionary context of different molecular machines. Bacteria and archaea are the biological subjects of our studies: their (relative) simplicity make them ideal subjects for study of basis biological principles, yet with considerable practical application in, for example, antibiotic development or sustainable re-utilization. To better understand evolution of molecular machinery we currently focus on a family of machines called type III secretion systems as a case study. Type III secretion systems function as nanoscale “3-D printers” to assemble one of two larger molecular machines: bacterial flagella and injectisomes. Bacterial flagella are nanoscale motors that spin helical filaments to act as a propellor for the bacterium, while injectisomes are nanoscale "syringes" that puncture host cells and pump toxic effector proteins into them. Nevertheless, the core type III secretion system found in both of flagella and injectisomes makes it clear that they are ancestrally related, making it an exciting case study to understand molecular evolution. We are also particularly interested in a number of curious variants of the flagellum that we recently identified, as these differences promise to shed light upon some basic principles of assembly, function, and evolution.
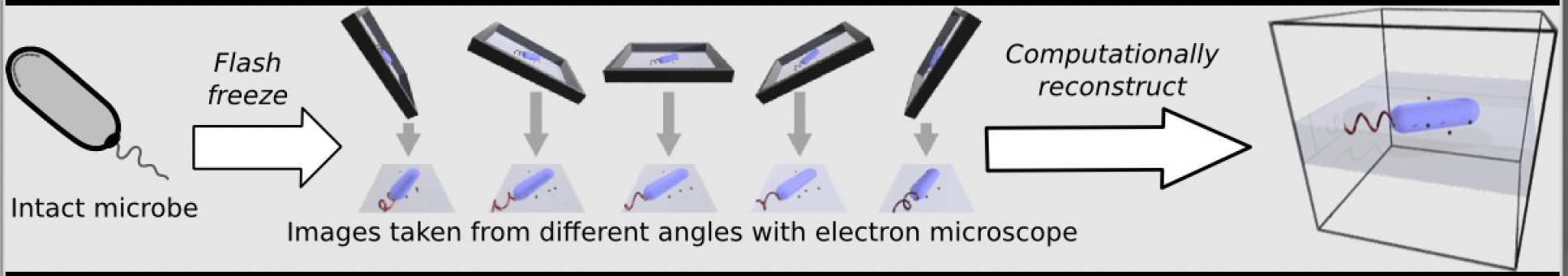 A core technique in the lab in electron cryo-tomography. This is where live biological samples (in our case, bacteria and archaea) are flash frozen onto special grids and imaged at multiple angles in the transmission electron microscope. We then use computational software to reconstruct the 3D shape of the object from the 2D pictures taken. The result is a 3 dimensional structure of the microscopic object we put in the microscope. Our current technologies allow us to "see" individual proteins and the structure of those proteins.
A core technique in the lab in electron cryo-tomography. This is where live biological samples (in our case, bacteria and archaea) are flash frozen onto special grids and imaged at multiple angles in the transmission electron microscope. We then use computational software to reconstruct the 3D shape of the object from the 2D pictures taken. The result is a 3 dimensional structure of the microscopic object we put in the microscope. Our current technologies allow us to "see" individual proteins and the structure of those proteins.
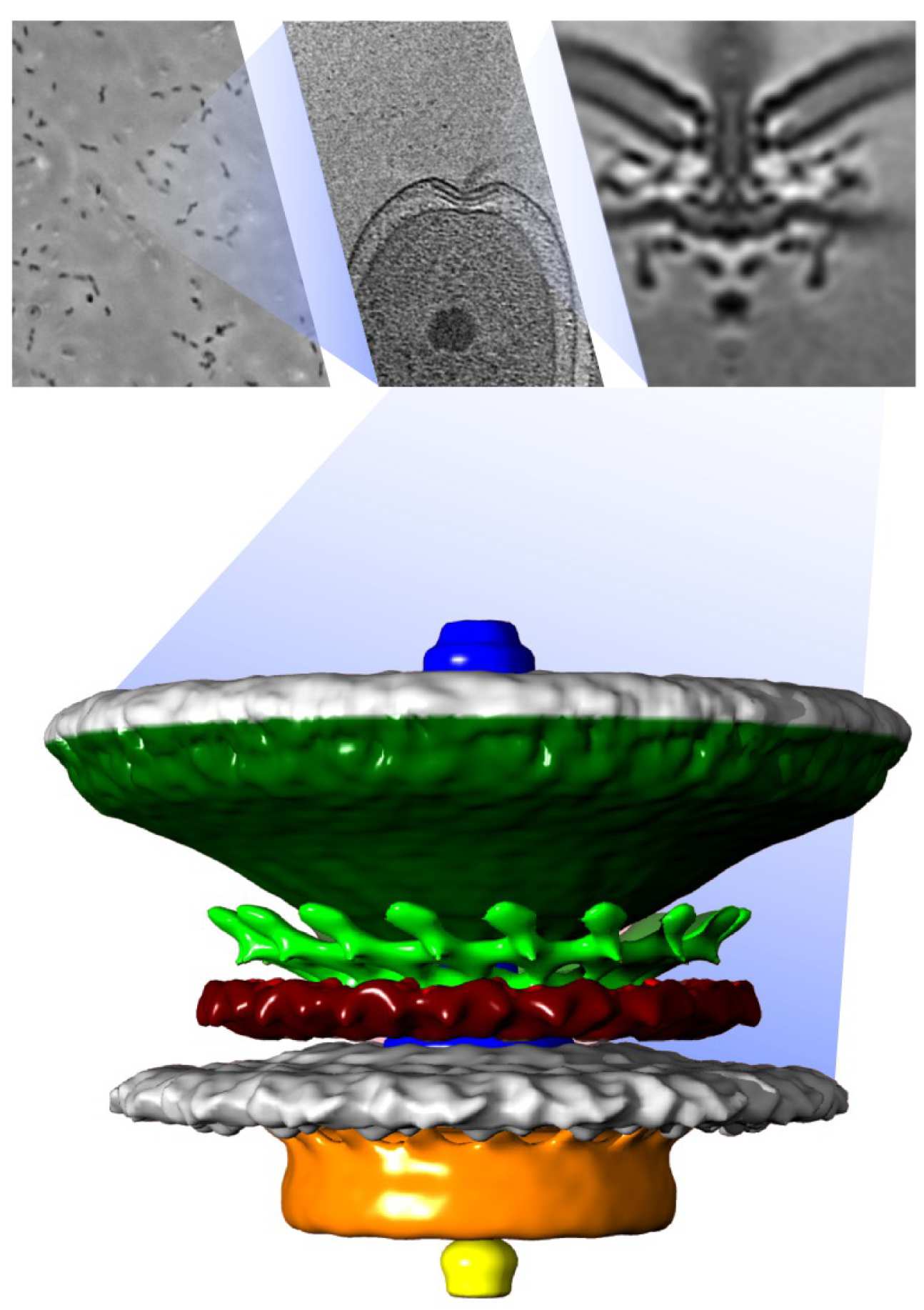
Together with the structures we can visualize, we also look at the evolutionary history of the molecular machines we study. To do this, we compare the gene and protein sequences that comprise the structure from many different species and look for similarities and differences. This type of analysis, called phylogenetic analysis, helps us understand when and how the parts of the structure evolved in the past to become what they are today.
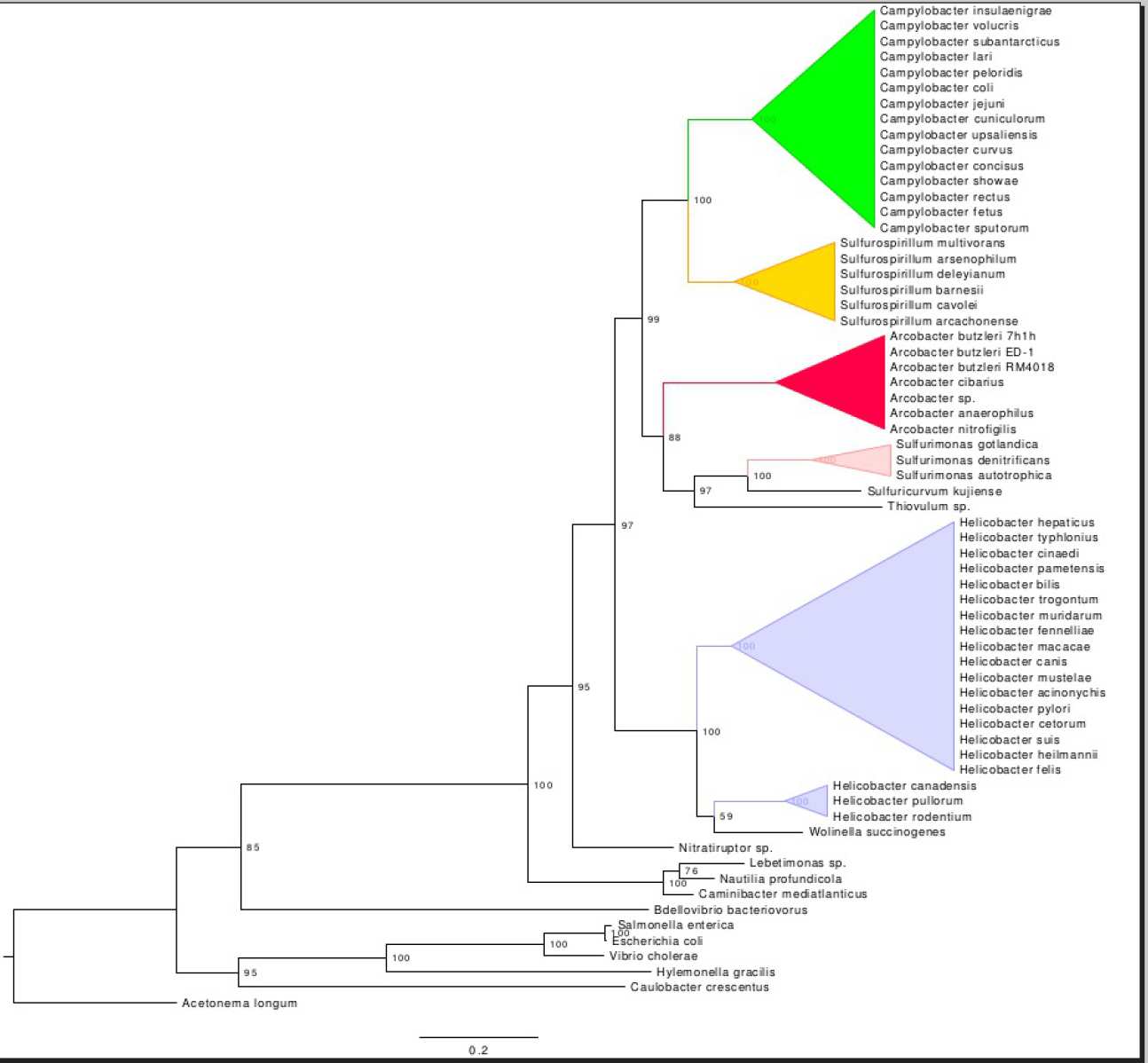
Bacteria and Archaea are our model organisms. Since the bacterial flagellar motor is one of the key molecular structures we study, looking at how organisms swim is an important characteristic to consider. Light microscopy and particle tracking are some of the ways we investigate this.
Tracking Salmonella enterica while swimming
Salmonella enterica swimming
Campylobacter jejuni tracked while swimming
Campylobacter jejuni swimming

Another area of active activity is finding ways to streamline high through-put data collection on the electron microscope.
Stay tuned as developments are implemented and showcased.